| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Peroxisome proliferator-activated receptor alpha |
|---|
| Ligand | BDBM28784 |
|---|
| Substrate/Competitor | BDBM10852 |
|---|
| Meas. Tech. | Cell-Based Transcription Assay |
|---|
| EC50 | 10.6±n/a nM |
|---|
| Citation |  Oon Han, H; Kim, SH; Kim, KH; Hur, GC; Joo Yim, H; Chung, HK; Ho Woo, S; Dong Koo, K; Lee, CS; Sung Koh, J; Kim, GT Design and synthesis of oxime ethers of alpha-acyl-beta-phenylpropanoic acids as PPAR dual agonists. Bioorg Med Chem Lett17:937-41 (2007) [PubMed] Article Oon Han, H; Kim, SH; Kim, KH; Hur, GC; Joo Yim, H; Chung, HK; Ho Woo, S; Dong Koo, K; Lee, CS; Sung Koh, J; Kim, GT Design and synthesis of oxime ethers of alpha-acyl-beta-phenylpropanoic acids as PPAR dual agonists. Bioorg Med Chem Lett17:937-41 (2007) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Solution Info, Assay Method |
|---|
| |
| Peroxisome proliferator-activated receptor alpha |
|---|
| Name: | Peroxisome proliferator-activated receptor alpha |
|---|
| Synonyms: | NR1C1 | Nuclear receptor subfamily 1 group C member 1 | PPAR | PPAR alpha/gamma | PPAR-alpha | PPARA | PPARA_HUMAN | Peroxisome Proliferator-Activated Receptor alpha | Peroxisome proliferator-activated receptor | Peroxisome proliferator-activated receptor alpha (PPAR alpha) |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 52222.08 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | Q07869 |
|---|
| Residue: | 468 |
|---|
| Sequence: | MVDTESPLCPLSPLEAGDLESPLSEEFLQEMGNIQEISQSIGEDSSGSFGFTEYQYLGSC
PGSDGSVITDTLSPASSPSSVTYPVVPGSVDESPSGALNIECRICGDKASGYHYGVHACE
GCKGFFRRTIRLKLVYDKCDRSCKIQKKNRNKCQYCRFHKCLSVGMSHNAIRFGRMPRSE
KAKLKAEILTCEHDIEDSETADLKSLAKRIYEAYLKNFNMNKVKARVILSGKASNNPPFV
IHDMETLCMAEKTLVAKLVANGIQNKEAEVRIFHCCQCTSVETVTELTEFAKAIPGFANL
DLNDQVTLLKYGVYEAIFAMLSSVMNKDGMLVAYGNGFITREFLKSLRKPFCDIMEPKFD
FAMKFNALELDDSDISLFVAAIICCGDRPGLLNVGHIEKMQEGIVHVLRLHLQSNHPDDI
FLFPKLLQKMADLRQLVTEHAQLVQIIKKTESDAALHPLLQEIYRDMY
|
|
|
|---|
| BDBM28784 |
|---|
| BDBM10852 |
|---|
| Name | BDBM28784 |
|---|
| Synonyms: | (3E)-3-{[(4-fluorophenyl)methoxy]imino}-2-({4-[2-(5-methyl-2-phenyl-1,3-oxazol-4-yl)ethoxy]phenyl}methyl)butanoic acid | alpha-acyl-beta-phenylpropanoic acid deriv., 11i |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C30H29FN2O5 |
|---|
| Mol. Mass. | 516.5601 |
|---|
| SMILES | C\C(=N/OCc1ccc(F)cc1)C(Cc1ccc(OCCc2nc(oc2C)-c2ccccc2)cc1)C(O)=O |
|---|
| Structure | 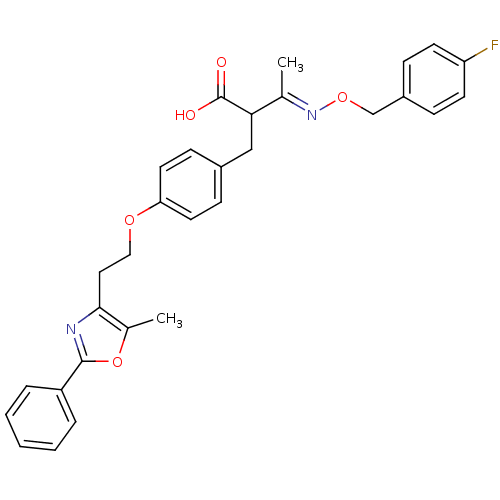
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Oon Han, H; Kim, SH; Kim, KH; Hur, GC; Joo Yim, H; Chung, HK; Ho Woo, S; Dong Koo, K; Lee, CS; Sung Koh, J; Kim, GT Design and synthesis of oxime ethers of alpha-acyl-beta-phenylpropanoic acids as PPAR dual agonists. Bioorg Med Chem Lett17:937-41 (2007) [PubMed] Article
Oon Han, H; Kim, SH; Kim, KH; Hur, GC; Joo Yim, H; Chung, HK; Ho Woo, S; Dong Koo, K; Lee, CS; Sung Koh, J; Kim, GT Design and synthesis of oxime ethers of alpha-acyl-beta-phenylpropanoic acids as PPAR dual agonists. Bioorg Med Chem Lett17:937-41 (2007) [PubMed] Article