| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Cannabinoid receptor 2 |
|---|
| Ligand | BDBM21278 |
|---|
| Substrate/Competitor | BDBM21244 |
|---|
| Meas. Tech. | CB1 Radioligand Binding Assay (Ki) and GTP-gamma-[35S] Binding Assay (EC50) |
|---|
| Ki | 1569±n/a nM |
|---|
| Citation |  Dow, RL; Carpino, PA; Hadcock, JR; Black, SC; Iredale, PA; DaSilva-Jardine, P; Schneider, SR; Paight, ES; Griffith, DA; Scott, DO; O'Connor, RE; Nduaka, CI Discovery of 2-(2-chlorophenyl)-3-(4-chlorophenyl)-7-(2,2-difluoropropyl)-6,7-dihydro-2H-pyrazolo[3,4-f][1,4]oxazepin-8(5H)-one (PF-514273), a novel, bicyclic lactam-based cannabinoid-1 receptor antagonist for the treatment of obesity. J Med Chem52:2652-5 (2009) [PubMed] Article Dow, RL; Carpino, PA; Hadcock, JR; Black, SC; Iredale, PA; DaSilva-Jardine, P; Schneider, SR; Paight, ES; Griffith, DA; Scott, DO; O'Connor, RE; Nduaka, CI Discovery of 2-(2-chlorophenyl)-3-(4-chlorophenyl)-7-(2,2-difluoropropyl)-6,7-dihydro-2H-pyrazolo[3,4-f][1,4]oxazepin-8(5H)-one (PF-514273), a novel, bicyclic lactam-based cannabinoid-1 receptor antagonist for the treatment of obesity. J Med Chem52:2652-5 (2009) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Solution Info, Assay Method |
|---|
| |
| Cannabinoid receptor 2 |
|---|
| Name: | Cannabinoid receptor 2 |
|---|
| Synonyms: | CANNABINOID CB2 | CB-2 | CB2 | CB2A | CB2B | CNR2 | CNR2_HUMAN | CX5 | Cannabinoid CB2 receptor | Cannabinoid receptor 2 (CB2) | Cannabinoid receptor 2 (CB2R) | hCB2 |
|---|
| Type: | G Protein-Coupled Receptor (GPCR) |
|---|
| Mol. Mass.: | 39690.94 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | P34972 |
|---|
| Residue: | 360 |
|---|
| Sequence: | MEECWVTEIANGSKDGLDSNPMKDYMILSGPQKTAVAVLCTLLGLLSALENVAVLYLILS
SHQLRRKPSYLFIGSLAGADFLASVVFACSFVNFHVFHGVDSKAVFLLKIGSVTMTFTAS
VGSLLLTAIDRYLCLRYPPSYKALLTRGRALVTLGIMWVLSALVSYLPLMGWTCCPRPCS
ELFPLIPNDYLLSWLLFIAFLFSGIIYTYGHVLWKAHQHVASLSGHQDRQVPGMARMRLD
VRLAKTLGLVLAVLLICWFPVLALMAHSLATTLSDQVKKAFAFCSMLCLINSMVNPVIYA
LRSGEIRSSAHHCLAHWKKCVRGLGSEAKEEAPRSSVTETEADGKITPWPDSRDLDLSDC
|
|
|
|---|
| BDBM21278 |
|---|
| BDBM21244 |
|---|
| Name | BDBM21278 |
|---|
| Synonyms: | 5-(4-chlorophenyl)-1-(2,4-dichlorophenyl)-4-methyl-N-(piperidin-1-yl)-1H-pyrazole-3-carboxamide | Acomplia | CHEMBL111 | CHEMBL558598 | RIMONABANT HYDROCHLORIDE | Rimonabant | SR 141716A | SR141716 | SR141716A | [3H]Rimonabant | [3H]SR141716A |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C22H21Cl3N4O |
|---|
| Mol. Mass. | 463.787 |
|---|
| SMILES | Cc1c(nn(c1-c1ccc(Cl)cc1)-c1ccc(Cl)cc1Cl)C(=O)NN1CCCCC1 |
|---|
| Structure | 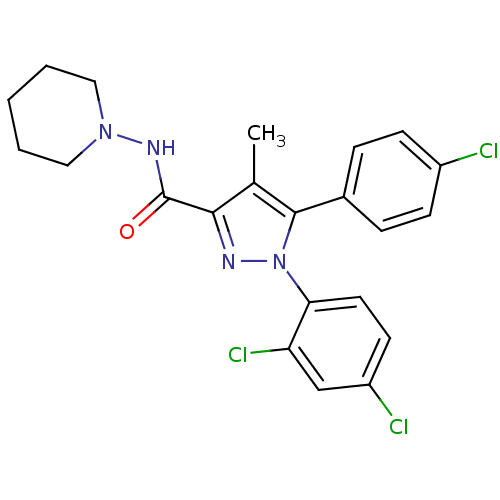
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Dow, RL; Carpino, PA; Hadcock, JR; Black, SC; Iredale, PA; DaSilva-Jardine, P; Schneider, SR; Paight, ES; Griffith, DA; Scott, DO; O'Connor, RE; Nduaka, CI Discovery of 2-(2-chlorophenyl)-3-(4-chlorophenyl)-7-(2,2-difluoropropyl)-6,7-dihydro-2H-pyrazolo[3,4-f][1,4]oxazepin-8(5H)-one (PF-514273), a novel, bicyclic lactam-based cannabinoid-1 receptor antagonist for the treatment of obesity. J Med Chem52:2652-5 (2009) [PubMed] Article
Dow, RL; Carpino, PA; Hadcock, JR; Black, SC; Iredale, PA; DaSilva-Jardine, P; Schneider, SR; Paight, ES; Griffith, DA; Scott, DO; O'Connor, RE; Nduaka, CI Discovery of 2-(2-chlorophenyl)-3-(4-chlorophenyl)-7-(2,2-difluoropropyl)-6,7-dihydro-2H-pyrazolo[3,4-f][1,4]oxazepin-8(5H)-one (PF-514273), a novel, bicyclic lactam-based cannabinoid-1 receptor antagonist for the treatment of obesity. J Med Chem52:2652-5 (2009) [PubMed] Article