| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Neprilysin |
|---|
| Ligand | BDBM50024098 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEBML_64511 |
|---|
| IC50 | 0.750000±n/a nM |
|---|
| Citation |  Fournié-Zaluski, MC; Lucas-Soroca, E; Devin, J; Roques, BP 1H NMR configurational correlation for retro-inverso dipeptides: application to the determination of the absolute configuration of"enkephalinase" inhibitors. Relationships between stereochemistry and enzyme recognition. J Med Chem29:751-7 (1986) [PubMed] Fournié-Zaluski, MC; Lucas-Soroca, E; Devin, J; Roques, BP 1H NMR configurational correlation for retro-inverso dipeptides: application to the determination of the absolute configuration of"enkephalinase" inhibitors. Relationships between stereochemistry and enzyme recognition. J Med Chem29:751-7 (1986) [PubMed] |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Neprilysin |
|---|
| Name: | Neprilysin |
|---|
| Synonyms: | Mme | NEP_RAT | Neprilysin | Neutral Endopeptidase (NEP) | Neutral endopeptidase 24.11 |
|---|
| Type: | Protein |
|---|
| Mol. Mass.: | 85789.59 |
|---|
| Organism: | Rattus norvegicus (Rat) |
|---|
| Description: | P07861 |
|---|
| Residue: | 750 |
|---|
| Sequence: | MGRSESQMDITDINAPKPKKKQRWTPLEISLSVLVLLLTIIAVTMIALYATYDDGICKSS
DCIKSAARLIQNMDASAEPCTDFFKYACGGWLKRNVIPETSSRYSNFDILRDELEVILKD
VLQEPKTEDIVAVQKAKTLYRSCINESAIDSRGGQPLLTLLPDIYGWPVASQNWEQTYGT
SWTAEKSIAQLNSKYGKKVLINFFVGTDDKNSTQHIIHFDQPRLGLPSRDYYECTGIYKE
ACTAYVDFMISVARLIRQEQRLPIDENQLSLEMNKVMELEKEIANATTKPEDRNDPMLLY
NKMTLAKLQNNFSLEINGKPFSWSNFTNEIMSTVNINIQNEEEVVVYAPEYLTKLKPILT
KYSPRDLQNLMSWRFIMDLVSSLSRNYKESRNAFRKALYGTTSETATWRRCANYVNGNME
NAVGRLYVEAAFAGESKHVVEDLIAQIREVFIQTLDDLTWMDAETKKKAEEKALAIKERI
GYPDDIISNENKLNNEYLELNYKEEEYFENIIQNLKFSQSKQLKKLREKVDKDEWISGAA
VVNAFYSSGRNQIVFPAGILQPPFFSARQSNSLNYGGIGMVIGHEITHGFDDNGRNFNKD
GDLVDWWTQQSANNFKDQSQCMVYQYGNFTWDLAGGQHLNGINTLGENIADNGGIGQAYR
AYQNYVKKNGEEKLLPGLDLNHKQLFFLNFAQVWCGTYRPEYAVNSIKTDVHSPGNFRII
GTLQNSAEFADAFHCRKNSYMNPERKCRVW
|
|
|
|---|
| BDBM50024098 |
|---|
| n/a |
|---|
| Name | BDBM50024098 |
|---|
| Synonyms: | (R,R) 2-(1-Mercaptomethyl-2-phenyl-ethylcarbamoyl)-4-methyl-pentanoic acid | CHEMBL164955 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C16H23NO3S |
|---|
| Mol. Mass. | 309.424 |
|---|
| SMILES | CC(C)C[C@H](C(O)=O)C(=O)N[C@H](CS)Cc1ccccc1 |
|---|
| Structure | 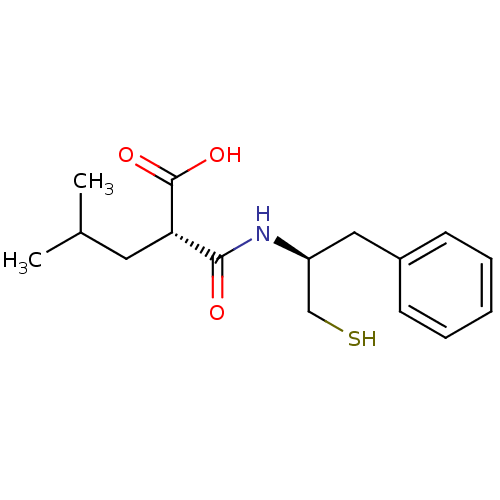
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Fournié-Zaluski, MC; Lucas-Soroca, E; Devin, J; Roques, BP 1H NMR configurational correlation for retro-inverso dipeptides: application to the determination of the absolute configuration of"enkephalinase" inhibitors. Relationships between stereochemistry and enzyme recognition. J Med Chem29:751-7 (1986) [PubMed]
Fournié-Zaluski, MC; Lucas-Soroca, E; Devin, J; Roques, BP 1H NMR configurational correlation for retro-inverso dipeptides: application to the determination of the absolute configuration of"enkephalinase" inhibitors. Relationships between stereochemistry and enzyme recognition. J Med Chem29:751-7 (1986) [PubMed]