

 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data
| Reaction Details | |||
|---|---|---|---|
 | Report a problem with these data | ||
| Target | Peroxisome proliferator-activated receptor gamma | ||
| Ligand | BDBM50519878 | ||
| Substrate/Competitor | n/a | ||
| Meas. Tech. | ChEMBL_1878653 (CHEMBL4380047) | ||
| IC50 | 78600±n/a nM | ||
| Citation |  Meijer, FA; Doveston, RG; de Vries, RMJM; Vos, GM; Vos, AAA; Leysen, S; Scheepstra, M; Ottmann, C; Milroy, LG; Brunsveld, L Ligand-Based Design of Allosteric Retinoic Acid Receptor-Related Orphan Receptor ?t (ROR?t) Inverse Agonists. J Med Chem63:241-259 (2020) [PubMed] Article Meijer, FA; Doveston, RG; de Vries, RMJM; Vos, GM; Vos, AAA; Leysen, S; Scheepstra, M; Ottmann, C; Milroy, LG; Brunsveld, L Ligand-Based Design of Allosteric Retinoic Acid Receptor-Related Orphan Receptor ?t (ROR?t) Inverse Agonists. J Med Chem63:241-259 (2020) [PubMed] Article | ||
| More Info.: | Get all data from this article, Assay Method | ||
| Peroxisome proliferator-activated receptor gamma | |||
| Name: | Peroxisome proliferator-activated receptor gamma | ||
| Synonyms: | NR1C3 | Nuclear receptor subfamily 1 group C member 3 | PPAR-gamma | PPARG | PPARG_HUMAN | Peroxisome proliferator-activated receptor | Peroxisome proliferator-activated receptor gamma (PPAR gamma) | Peroxisome proliferator-activated receptor gamma (PPARG) | Peroxisome proliferator-activated receptor gamma (PPARγ) | Peroxisome proliferator-activated receptor gamma/Nuclear receptor corepressor 2 | peroxisome proliferator-activated receptor gamma isoform 2 | ||
| Type: | Nuclear Receptor | ||
| Mol. Mass.: | 57613.46 | ||
| Organism: | Homo sapiens (Human) | ||
| Description: | P37231 | ||
| Residue: | 505 | ||
| Sequence: |
| ||
| BDBM50519878 | |||
| n/a | |||
| Name | BDBM50519878 | ||
| Synonyms: | CHEMBL4550831 | ||
| Type | Small organic molecule | ||
| Emp. Form. | C24H16ClF3N2O3 | ||
| Mol. Mass. | 472.844 | ||
| SMILES | OC(=O)c1ccc(NCc2c(noc2-c2ccccc2)-c2c(Cl)cccc2C(F)(F)F)cc1 |(17.9,-10.08,;17.57,-8.58,;18.7,-7.54,;16.1,-8.11,;14.96,-9.15,;13.5,-8.68,;13.17,-7.18,;11.71,-6.71,;10.57,-7.75,;9.11,-7.28,;7.86,-8.2,;6.61,-7.29,;7.07,-5.82,;8.62,-5.81,;9.51,-4.57,;11.05,-4.73,;11.94,-3.48,;11.31,-2.07,;9.77,-1.93,;8.88,-3.17,;7.88,-9.78,;9.25,-10.55,;10.61,-9.73,;9.28,-12.12,;7.92,-12.93,;6.54,-12.16,;6.52,-10.58,;5.15,-9.81,;5.13,-8.27,;3.83,-10.59,;3.81,-9.03,;14.3,-6.14,;15.77,-6.6,)| | ||
| Structure | 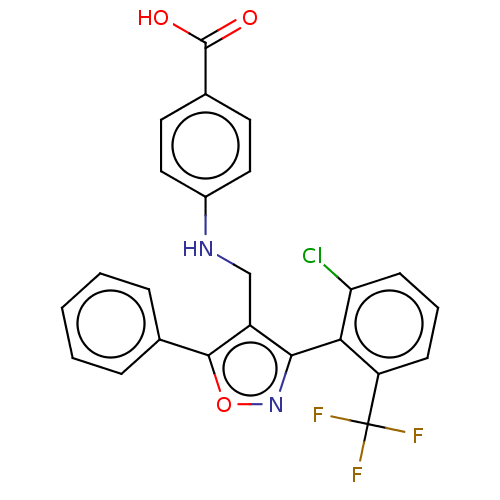
| ||