| Reaction Details |
|---|
 | Report a problem with these data |
| Target | RAC-beta serine/threonine-protein kinase |
|---|
| Ligand | BDBM50536168 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_1933005 (CHEMBL4478657) |
|---|
| IC50 | 5.4±n/a nM |
|---|
| Citation |  Lapierre, JM; Eathiraj, S; Vensel, D; Liu, Y; Bull, CO; Cornell-Kennon, S; Iimura, S; Kelleher, EW; Kizer, DE; Koerner, S; Makhija, S; Matsuda, A; Moussa, M; Namdev, N; Savage, RE; Szwaya, J; Volckova, E; Westlund, N; Wu, H; Schwartz, B Discovery of 3-(3-(4-(1-Aminocyclobutyl)phenyl)-5-phenyl-3H-imidazo[4,5-b]pyridin-2-yl)pyridin-2-amine (ARQ 092): An Orally Bioavailable, Selective, and Potent Allosteric AKT Inhibitor. J Med Chem59:6455-69 (2016) [PubMed] Article Lapierre, JM; Eathiraj, S; Vensel, D; Liu, Y; Bull, CO; Cornell-Kennon, S; Iimura, S; Kelleher, EW; Kizer, DE; Koerner, S; Makhija, S; Matsuda, A; Moussa, M; Namdev, N; Savage, RE; Szwaya, J; Volckova, E; Westlund, N; Wu, H; Schwartz, B Discovery of 3-(3-(4-(1-Aminocyclobutyl)phenyl)-5-phenyl-3H-imidazo[4,5-b]pyridin-2-yl)pyridin-2-amine (ARQ 092): An Orally Bioavailable, Selective, and Potent Allosteric AKT Inhibitor. J Med Chem59:6455-69 (2016) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| RAC-beta serine/threonine-protein kinase |
|---|
| Name: | RAC-beta serine/threonine-protein kinase |
|---|
| Synonyms: | AKT2 | AKT2_HUMAN | PKB beta | Protein kinase Akt-2 | Protein kinase B (Akt 2) | Protein kinase B beta (AKT2) | Protein kinase B, beta | RAC-PK-beta | RAC-alpha serine/threonine-protein kinase (AKT2) | RAC-beta serine/threonine-protein kinase | RAC-beta serine/threonine-protein kinase (AKT2) | Serine/threonine-protein kinase AKT | Serine/threonine-protein kinase AKT2 | Serine/threonine-protein kinase Rac alpha/beta |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 55766.64 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | n/a |
|---|
| Residue: | 481 |
|---|
| Sequence: | MNEVSVIKEGWLHKRGEYIKTWRPRYFLLKSDGSFIGYKERPEAPDQTLPPLNNFSVAEC
QLMKTERPRPNTFVIRCLQWTTVIERTFHVDSPDEREEWMRAIQMVANSLKQRAPGEDPM
DYKCGSPSDSSTTEEMEVAVSKARAKVTMNDFDYLKLLGKGTFGKVILVREKATGRYYAM
KILRKEVIIAKDEVAHTVTESRVLQNTRHPFLTALKYAFQTHDRLCFVMEYANGGELFFH
LSRERVFTEERARFYGAEIVSALEYLHSRDVVYRDIKLENLMLDKDGHIKITDFGLCKEG
ISDGATMKTFCGTPEYLAPEVLEDNDYGRAVDWWGLGVVMYEMMCGRLPFYNQDHERLFE
LILMEEIRFPRTLSPEAKSLLAGLLKKDPKQRLGGGPSDAKEVMEHRFFLSINWQDVVQK
KLLPPFKPQVTSEVDTRYFDDEFTAQSITITPPDRYDSLGLLELDQRTHFPQFSYSASIR
E
|
|
|
|---|
| BDBM50536168 |
|---|
| n/a |
|---|
| Name | BDBM50536168 |
|---|
| Synonyms: | CHEMBL4574111 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C27H26ClN7 |
|---|
| Mol. Mass. | 483.995 |
|---|
| SMILES | Cl.Nc1cccc(c1)-c1ccc2nc(-c3cccnc3N)n(-c3ccc(cc3)C3(N)CCC3)c2n1 |
|---|
| Structure | 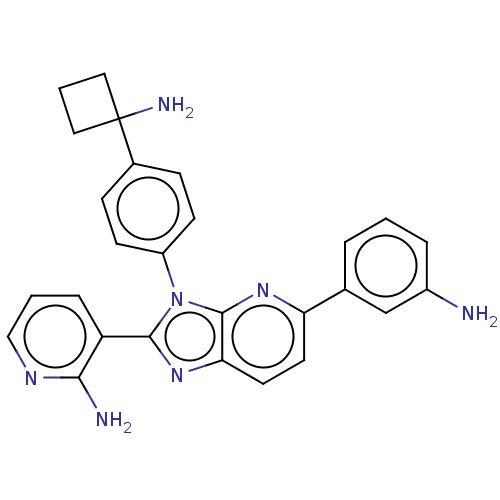
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Lapierre, JM; Eathiraj, S; Vensel, D; Liu, Y; Bull, CO; Cornell-Kennon, S; Iimura, S; Kelleher, EW; Kizer, DE; Koerner, S; Makhija, S; Matsuda, A; Moussa, M; Namdev, N; Savage, RE; Szwaya, J; Volckova, E; Westlund, N; Wu, H; Schwartz, B Discovery of 3-(3-(4-(1-Aminocyclobutyl)phenyl)-5-phenyl-3H-imidazo[4,5-b]pyridin-2-yl)pyridin-2-amine (ARQ 092): An Orally Bioavailable, Selective, and Potent Allosteric AKT Inhibitor. J Med Chem59:6455-69 (2016) [PubMed] Article
Lapierre, JM; Eathiraj, S; Vensel, D; Liu, Y; Bull, CO; Cornell-Kennon, S; Iimura, S; Kelleher, EW; Kizer, DE; Koerner, S; Makhija, S; Matsuda, A; Moussa, M; Namdev, N; Savage, RE; Szwaya, J; Volckova, E; Westlund, N; Wu, H; Schwartz, B Discovery of 3-(3-(4-(1-Aminocyclobutyl)phenyl)-5-phenyl-3H-imidazo[4,5-b]pyridin-2-yl)pyridin-2-amine (ARQ 092): An Orally Bioavailable, Selective, and Potent Allosteric AKT Inhibitor. J Med Chem59:6455-69 (2016) [PubMed] Article