

 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data
| Reaction Details | |||
|---|---|---|---|
 | Report a problem with these data | ||
| Target | Tyrosine-protein kinase Lck | ||
| Ligand | BDBM50132391 | ||
| Substrate/Competitor | n/a | ||
| Meas. Tech. | ChEBML_221488 | ||
| IC50 | 1400±n/a nM | ||
| Citation |  He, W; Myers, MR; Hanney, B; Spada, AP; Bilder, G; Galzcinski, H; Amin, D; Needle, S; Page, K; Jayyosi, Z; Perrone, MH Potent quinoxaline-based inhibitors of PDGF receptor tyrosine kinase activity. Part 2: the synthesis and biological activities of RPR127963 an orally bioavailable inhibitor. Bioorg Med Chem Lett13:3097-100 (2003) [PubMed] He, W; Myers, MR; Hanney, B; Spada, AP; Bilder, G; Galzcinski, H; Amin, D; Needle, S; Page, K; Jayyosi, Z; Perrone, MH Potent quinoxaline-based inhibitors of PDGF receptor tyrosine kinase activity. Part 2: the synthesis and biological activities of RPR127963 an orally bioavailable inhibitor. Bioorg Med Chem Lett13:3097-100 (2003) [PubMed] | ||
| More Info.: | Get all data from this article, Assay Method | ||
| Tyrosine-protein kinase Lck | |||
| Name: | Tyrosine-protein kinase Lck | ||
| Synonyms: | 2.7.10.2 | LCK | LCK_HUMAN | LSK | Leukocyte C-terminal Src kinase | Lymphocyte cell-specific protein-tyrosine kinase | Lymphocyte-specific protein tyrosine kinase | P56-LCK | Protein YT16 | Proto-oncogene Lck | Proto-oncogene tyrosine-protein kinase LCK | Src/Lck kinase | T cell-specific protein-tyrosine kinase | ||
| Type: | n/a | ||
| Mol. Mass.: | 57987.83 | ||
| Organism: | Homo sapiens (Human) | ||
| Description: | P06239 | ||
| Residue: | 509 | ||
| Sequence: |
| ||
| BDBM50132391 | |||
| n/a | |||
| Name | BDBM50132391 | ||
| Synonyms: | 4-((R)-6,7-Dimethoxy-quinoxalin-2-ylamino)-cyclohexanol | 4-(6,7-Dimethoxy-quinoxalin-2-ylamino)-cyclohexanol | CHEMBL104466 | RP-127963 | ||
| Type | Small organic molecule | ||
| Emp. Form. | C16H21N3O3 | ||
| Mol. Mass. | 303.3562 | ||
| SMILES | COc1cc2ncc(N[C@H]3CC[C@H](O)CC3)nc2cc1OC |wU:12.12,wD:9.8,(-.13,-6.56,;-.13,-5.02,;1.2,-4.25,;2.53,-5.02,;3.88,-4.23,;5.21,-5,;6.54,-4.23,;6.54,-2.69,;7.85,-1.9,;9.2,-2.67,;9.2,-4.21,;10.53,-4.98,;11.86,-4.21,;13.19,-4.98,;11.86,-2.67,;10.53,-1.9,;5.19,-1.92,;3.88,-2.69,;2.53,-1.92,;1.2,-2.71,;-.13,-1.94,;-.13,-.4,)| | ||
| Structure | 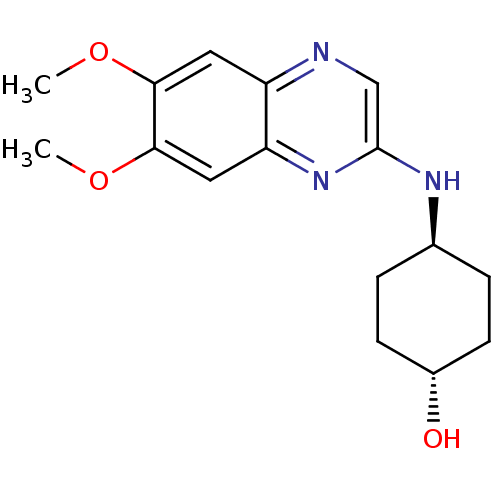
| ||