

 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data
| Reaction Details | |||
|---|---|---|---|
 | Report a problem with these data | ||
| Target | Leucine-rich repeat serine/threonine-protein kinase 2 | ||
| Ligand | BDBM50590220 | ||
| Substrate/Competitor | n/a | ||
| Meas. Tech. | ChEMBL_2195620 (CHEMBL5108136) | ||
| IC50 | 2.0±n/a nM | ||
| Citation |  Gulati, A; Yeung, CS; Lapointe, B; Kattar, SD; Gunaydin, H; Scott, JD; Childers, KK; Methot, JL; Simov, V; Kurukulasuriya, R; Pio, B; Morriello, GJ; Liu, P; Tang, H; Neelamkavil, S; Wood, HB; Rada, VL; Ardolino, MJ; Yan, XC; Palte, R; Otte, K; Faltus, R; Woodhouse, J; Hegde, LG; Ciaccio, P; Minnihan, EC; DiMauro, EF; Fell, MJ; Fuller, PH; Ellis, JM Optimization of brain-penetrant picolinamide derived leucine-rich repeat kinase 2 (LRRK2) inhibitors. RSC Med Chem12:1164-1173 (2021) [PubMed] Article Gulati, A; Yeung, CS; Lapointe, B; Kattar, SD; Gunaydin, H; Scott, JD; Childers, KK; Methot, JL; Simov, V; Kurukulasuriya, R; Pio, B; Morriello, GJ; Liu, P; Tang, H; Neelamkavil, S; Wood, HB; Rada, VL; Ardolino, MJ; Yan, XC; Palte, R; Otte, K; Faltus, R; Woodhouse, J; Hegde, LG; Ciaccio, P; Minnihan, EC; DiMauro, EF; Fell, MJ; Fuller, PH; Ellis, JM Optimization of brain-penetrant picolinamide derived leucine-rich repeat kinase 2 (LRRK2) inhibitors. RSC Med Chem12:1164-1173 (2021) [PubMed] Article | ||
| More Info.: | Get all data from this article, Assay Method | ||
| Leucine-rich repeat serine/threonine-protein kinase 2 | |||
| Name: | Leucine-rich repeat serine/threonine-protein kinase 2 | ||
| Synonyms: | Dardarin | LRRK2 | LRRK2_HUMAN | Leucine-Rich Repeat Kinase 2 Protein (LRRK2) | Leucine-rich repeat serine/threonine-protein kinase 2 | Leucine-rich repeat serine/threonine-protein kinase 2 (LRRK2) | PARK8 | ||
| Type: | Protein | ||
| Mol. Mass.: | 286113.20 | ||
| Organism: | Homo sapiens (Human) | ||
| Description: | Q5S007 | ||
| Residue: | 2527 | ||
| Sequence: |
| ||
| BDBM50590220 | |||
| n/a | |||
| Name | BDBM50590220 | ||
| Synonyms: | CHEMBL5202285 | ||
| Type | Small organic molecule | ||
| Emp. Form. | C24H25F3N6O2 | ||
| Mol. Mass. | 486.4895 | ||
| SMILES | Cn1cc(cn1)-c1cccc(n1)C(=O)Nc1cc(ncc1C(F)(F)F)N1CC2(CC1C2)C(C)(C)O |(6.51,-7.33,;5.17,-6.56,;5.17,-4.99,;3.62,-4.59,;2.8,-5.89,;3.77,-7.1,;3.22,-3.11,;4.3,-2.02,;3.91,-.53,;2.42,-.13,;1.33,-1.22,;1.73,-2.72,;-.15,-.82,;-.55,.66,;-1.24,-1.91,;-2.73,-1.51,;-3.13,-.02,;-4.62,.37,;-5.7,-.72,;-5.3,-2.2,;-3.82,-2.6,;-3.42,-4.09,;-3.02,-5.58,;-4.51,-5.18,;-1.93,-4.49,;-5.02,1.86,;-4.2,3.15,;-5.16,4.36,;-6.51,3.83,;-6.51,2.25,;-5.37,2.95,;-4.76,5.85,;-5.85,6.94,;-3.28,6.24,;-4.37,7.33,)| | ||
| Structure | 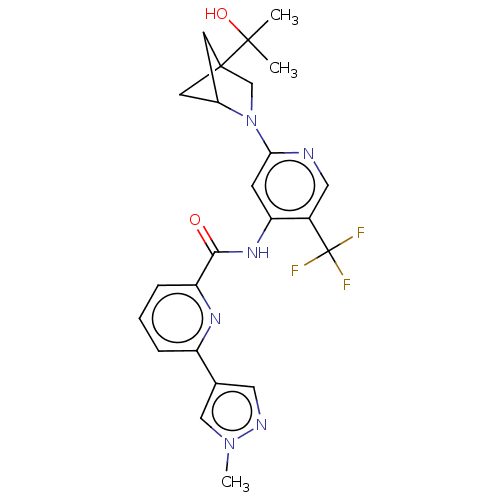
| ||