| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Beta-secretase 1 |
|---|
| Ligand | BDBM50457586 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_2266459 |
|---|
| IC50 | 59±n/a nM |
|---|
| Citation |  Oguma, T; Uehara, S; Nakahara, K; Okuyama, Y; Fuchino, K; Suzuki, N; Kan, Y; Kanegawa, N; Ogata, Y; Kusakabe, KI A Quantum Mechanics-Based Method to Predict Intramolecular Hydrogen Bond Formation Reflecting P-glycoprotein Recognition. ACS Med Chem Lett14:223-228 (2023) [PubMed] Oguma, T; Uehara, S; Nakahara, K; Okuyama, Y; Fuchino, K; Suzuki, N; Kan, Y; Kanegawa, N; Ogata, Y; Kusakabe, KI A Quantum Mechanics-Based Method to Predict Intramolecular Hydrogen Bond Formation Reflecting P-glycoprotein Recognition. ACS Med Chem Lett14:223-228 (2023) [PubMed] |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Beta-secretase 1 |
|---|
| Name: | Beta-secretase 1 |
|---|
| Synonyms: | ASP2 | Asp 2 | Aspartyl protease 2 | BACE | BACE1 | BACE1_HUMAN | Beta-secretase (BACE) | Beta-secretase 1 | Beta-secretase 1 (BACE 1) | Beta-secretase 1 (BACE-1) | Beta-secretase 1 (BACE1) | Beta-site APP cleaving enzyme 1 | Beta-site amyloid precursor protein cleaving enzyme 1 | KIAA1149 | Memapsin-2 | Membrane-associated aspartic protease 2 | beta-Secretase (BACE-1) | beta-Secretase (BACE1) |
|---|
| Type: | Protein |
|---|
| Mol. Mass.: | 55755.10 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | P56817 |
|---|
| Residue: | 501 |
|---|
| Sequence: | MAQALPWLLLWMGAGVLPAHGTQHGIRLPLRSGLGGAPLGLRLPRETDEEPEEPGRRGSF
VEMVDNLRGKSGQGYYVEMTVGSPPQTLNILVDTGSSNFAVGAAPHPFLHRYYQRQLSST
YRDLRKGVYVPYTQGKWEGELGTDLVSIPHGPNVTVRANIAAITESDKFFINGSNWEGIL
GLAYAEIARPDDSLEPFFDSLVKQTHVPNLFSLQLCGAGFPLNQSEVLASVGGSMIIGGI
DHSLYTGSLWYTPIRREWYYEVIIVRVEINGQDLKMDCKEYNYDKSIVDSGTTNLRLPKK
VFEAAVKSIKAASSTEKFPDGFWLGEQLVCWQAGTTPWNIFPVISLYLMGEVTNQSFRIT
ILPQQYLRPVEDVATSQDDCYKFAISQSSTGTVMGAVIMEGFYVVFDRARKRIGFAVSAC
HVHDEFRTAAVEGPFVTLDMEDCGYNIPQTDESTLMTIAYVMAAICALFMLPLCLMVCQW
RCLRCLRQQHDDFADDISLLK
|
|
|
|---|
| BDBM50457586 |
|---|
| n/a |
|---|
| Name | BDBM50457586 |
|---|
| Synonyms: | CHEMBL4213486 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C19H15F2N5O2 |
|---|
| Mol. Mass. | 383.3515 |
|---|
| SMILES | C[C@]1(C=C(CF)OC(N)=N1)c1cc(NC(=O)c2ccc(cn2)C#N)ccc1F |r,c:8,t:2| |
|---|
| Structure | 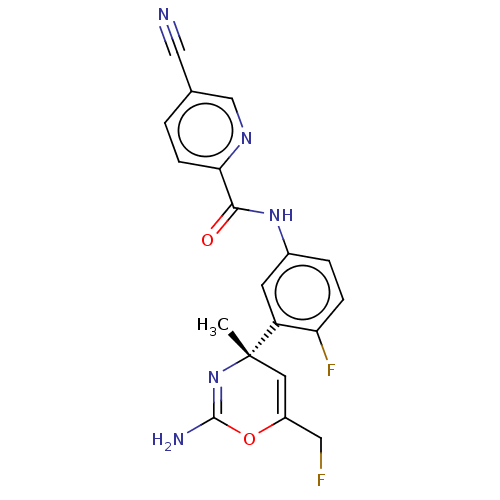
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Oguma, T; Uehara, S; Nakahara, K; Okuyama, Y; Fuchino, K; Suzuki, N; Kan, Y; Kanegawa, N; Ogata, Y; Kusakabe, KI A Quantum Mechanics-Based Method to Predict Intramolecular Hydrogen Bond Formation Reflecting P-glycoprotein Recognition. ACS Med Chem Lett14:223-228 (2023) [PubMed]
Oguma, T; Uehara, S; Nakahara, K; Okuyama, Y; Fuchino, K; Suzuki, N; Kan, Y; Kanegawa, N; Ogata, Y; Kusakabe, KI A Quantum Mechanics-Based Method to Predict Intramolecular Hydrogen Bond Formation Reflecting P-glycoprotein Recognition. ACS Med Chem Lett14:223-228 (2023) [PubMed]