| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Peroxisome proliferator-activated receptor alpha |
|---|
| Ligand | BDBM50195708 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_425035 (CHEMBL912556) |
|---|
| EC50 | 986±n/a nM |
|---|
| Citation |  Warshawsky, AM; Alt, CA; Brozinick, JT; Harkness, AR; Hawkins, ED; Henry, JR; Matthews, DP; Miller, AR; Misener, EA; Montrose-Rafizadeh, C; Rhodes, GA; Shen, Q; Vance, JA; Udodong, UE; Wang, M; Zhang, TY; Zink, RW Synthesis and evaluation of aminomethyl dihydrocinnamates as a new class of PPAR ligands. Bioorg Med Chem Lett16:6328-33 (2006) [PubMed] Article Warshawsky, AM; Alt, CA; Brozinick, JT; Harkness, AR; Hawkins, ED; Henry, JR; Matthews, DP; Miller, AR; Misener, EA; Montrose-Rafizadeh, C; Rhodes, GA; Shen, Q; Vance, JA; Udodong, UE; Wang, M; Zhang, TY; Zink, RW Synthesis and evaluation of aminomethyl dihydrocinnamates as a new class of PPAR ligands. Bioorg Med Chem Lett16:6328-33 (2006) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Peroxisome proliferator-activated receptor alpha |
|---|
| Name: | Peroxisome proliferator-activated receptor alpha |
|---|
| Synonyms: | NR1C1 | Nuclear receptor subfamily 1 group C member 1 | PPAR | PPAR alpha/gamma | PPAR-alpha | PPARA | PPARA_HUMAN | Peroxisome Proliferator-Activated Receptor alpha | Peroxisome proliferator-activated receptor | Peroxisome proliferator-activated receptor alpha (PPAR alpha) |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 52222.08 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | Q07869 |
|---|
| Residue: | 468 |
|---|
| Sequence: | MVDTESPLCPLSPLEAGDLESPLSEEFLQEMGNIQEISQSIGEDSSGSFGFTEYQYLGSC
PGSDGSVITDTLSPASSPSSVTYPVVPGSVDESPSGALNIECRICGDKASGYHYGVHACE
GCKGFFRRTIRLKLVYDKCDRSCKIQKKNRNKCQYCRFHKCLSVGMSHNAIRFGRMPRSE
KAKLKAEILTCEHDIEDSETADLKSLAKRIYEAYLKNFNMNKVKARVILSGKASNNPPFV
IHDMETLCMAEKTLVAKLVANGIQNKEAEVRIFHCCQCTSVETVTELTEFAKAIPGFANL
DLNDQVTLLKYGVYEAIFAMLSSVMNKDGMLVAYGNGFITREFLKSLRKPFCDIMEPKFD
FAMKFNALELDDSDISLFVAAIICCGDRPGLLNVGHIEKMQEGIVHVLRLHLQSNHPDDI
FLFPKLLQKMADLRQLVTEHAQLVQIIKKTESDAALHPLLQEIYRDMY
|
|
|
|---|
| BDBM50195708 |
|---|
| n/a |
|---|
| Name | BDBM50195708 |
|---|
| Synonyms: | 3-(2-((isopropoxycarbonyl)methyl)-4-(2-(5-methyl-2-morpholinothiazol-4-yl)ethoxy)phenyl)propanoic acid | CHEMBL223351 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C24H33N3O6S |
|---|
| Mol. Mass. | 491.6 |
|---|
| SMILES | CC(C)OC(=O)NCc1cc(OCCc2nc(sc2C)N2CCOCC2)ccc1CCC(O)=O |
|---|
| Structure | 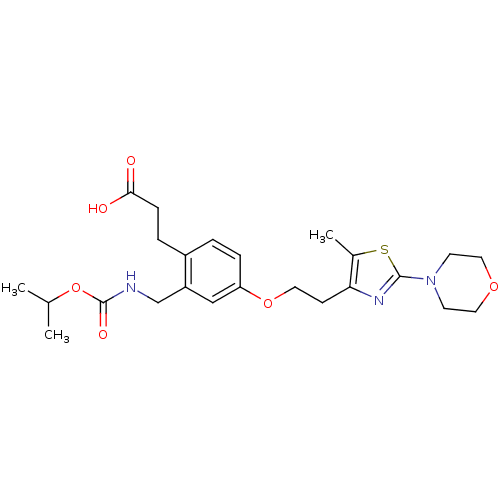
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Warshawsky, AM; Alt, CA; Brozinick, JT; Harkness, AR; Hawkins, ED; Henry, JR; Matthews, DP; Miller, AR; Misener, EA; Montrose-Rafizadeh, C; Rhodes, GA; Shen, Q; Vance, JA; Udodong, UE; Wang, M; Zhang, TY; Zink, RW Synthesis and evaluation of aminomethyl dihydrocinnamates as a new class of PPAR ligands. Bioorg Med Chem Lett16:6328-33 (2006) [PubMed] Article
Warshawsky, AM; Alt, CA; Brozinick, JT; Harkness, AR; Hawkins, ED; Henry, JR; Matthews, DP; Miller, AR; Misener, EA; Montrose-Rafizadeh, C; Rhodes, GA; Shen, Q; Vance, JA; Udodong, UE; Wang, M; Zhang, TY; Zink, RW Synthesis and evaluation of aminomethyl dihydrocinnamates as a new class of PPAR ligands. Bioorg Med Chem Lett16:6328-33 (2006) [PubMed] Article