| Citation |  Xiong, Y; Teegarden, BR; Choi, JS; Strah-Pleynet, S; Decaire, M; Jayakumar, H; Dosa, PI; Casper, MD; Pham, L; Feichtinger, K; Ullman, B; Adams, J; Yuskin, D; Frazer, J; Morgan, M; Sadeque, A; Chen, W; Webb, RR; Connolly, DT; Semple, G; Al-Shamma, H Discovery and structure-activity relationship of 3-methoxy-N-(3-(1-methyl-1H-pyrazol-5-yl)-4-(2-morpholinoethoxy)phenyl)benzamide (APD791): a highly selective 5-hydroxytryptamine2A receptor inverse agonist for the treatment of arterial thrombosis. J Med Chem53:4412-21 (2010) [PubMed] Article Xiong, Y; Teegarden, BR; Choi, JS; Strah-Pleynet, S; Decaire, M; Jayakumar, H; Dosa, PI; Casper, MD; Pham, L; Feichtinger, K; Ullman, B; Adams, J; Yuskin, D; Frazer, J; Morgan, M; Sadeque, A; Chen, W; Webb, RR; Connolly, DT; Semple, G; Al-Shamma, H Discovery and structure-activity relationship of 3-methoxy-N-(3-(1-methyl-1H-pyrazol-5-yl)-4-(2-morpholinoethoxy)phenyl)benzamide (APD791): a highly selective 5-hydroxytryptamine2A receptor inverse agonist for the treatment of arterial thrombosis. J Med Chem53:4412-21 (2010) [PubMed] Article |
|---|
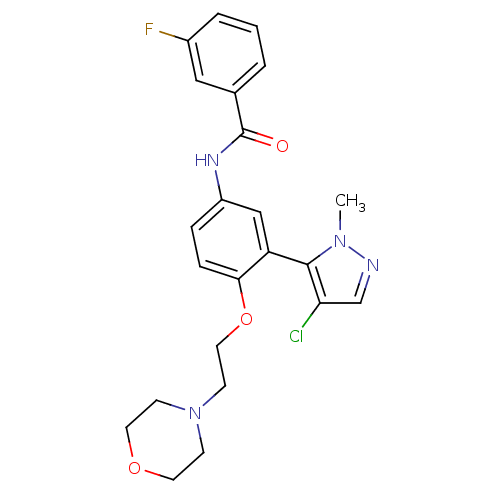
| SMILES | Cn1ncc(Cl)c1-c1cc(NC(=O)c2cccc(F)c2)ccc1OCCN1CCOCC1 |(-5.98,-34.34,;-5.66,-35.84,;-6.69,-36.99,;-5.92,-38.32,;-4.41,-38,;-3.26,-39.03,;-4.25,-36.47,;-2.92,-35.7,;-1.59,-36.48,;-.25,-35.7,;1.09,-36.47,;2.42,-35.7,;2.42,-34.16,;3.75,-36.47,;3.75,-38,;5.08,-38.77,;6.41,-38,;6.41,-36.45,;7.74,-35.67,;5.07,-35.69,;-.26,-34.15,;-1.59,-33.39,;-2.92,-34.16,;-4.25,-33.39,;-4.25,-31.85,;-5.59,-31.08,;-5.59,-29.54,;-4.25,-28.77,;-4.25,-27.24,;-5.58,-26.46,;-6.92,-27.23,;-6.92,-28.77,)| |
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Xiong, Y; Teegarden, BR; Choi, JS; Strah-Pleynet, S; Decaire, M; Jayakumar, H; Dosa, PI; Casper, MD; Pham, L; Feichtinger, K; Ullman, B; Adams, J; Yuskin, D; Frazer, J; Morgan, M; Sadeque, A; Chen, W; Webb, RR; Connolly, DT; Semple, G; Al-Shamma, H Discovery and structure-activity relationship of 3-methoxy-N-(3-(1-methyl-1H-pyrazol-5-yl)-4-(2-morpholinoethoxy)phenyl)benzamide (APD791): a highly selective 5-hydroxytryptamine2A receptor inverse agonist for the treatment of arterial thrombosis. J Med Chem53:4412-21 (2010) [PubMed] Article
Xiong, Y; Teegarden, BR; Choi, JS; Strah-Pleynet, S; Decaire, M; Jayakumar, H; Dosa, PI; Casper, MD; Pham, L; Feichtinger, K; Ullman, B; Adams, J; Yuskin, D; Frazer, J; Morgan, M; Sadeque, A; Chen, W; Webb, RR; Connolly, DT; Semple, G; Al-Shamma, H Discovery and structure-activity relationship of 3-methoxy-N-(3-(1-methyl-1H-pyrazol-5-yl)-4-(2-morpholinoethoxy)phenyl)benzamide (APD791): a highly selective 5-hydroxytryptamine2A receptor inverse agonist for the treatment of arterial thrombosis. J Med Chem53:4412-21 (2010) [PubMed] Article