| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Peroxisome proliferator-activated receptor delta |
|---|
| Ligand | BDBM28701 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_700929 (CHEMBL1648428) |
|---|
| EC50 | 18900±n/a nM |
|---|
| Citation |  Makadia, P; Shah, SR; Pingali, H; Zaware, P; Patel, D; Pola, S; Thube, B; Priyadarshini, P; Suthar, D; Shah, M; Giri, S; Trivedi, C; Jain, M; Patel, P; Bahekar, R Effect of structurally constrained oxime-ether linker on PPAR subtype selectivity: Discovery of a novel and potent series of PPAR-pan agonists. Bioorg Med Chem19:771-82 (2011) [PubMed] Article Makadia, P; Shah, SR; Pingali, H; Zaware, P; Patel, D; Pola, S; Thube, B; Priyadarshini, P; Suthar, D; Shah, M; Giri, S; Trivedi, C; Jain, M; Patel, P; Bahekar, R Effect of structurally constrained oxime-ether linker on PPAR subtype selectivity: Discovery of a novel and potent series of PPAR-pan agonists. Bioorg Med Chem19:771-82 (2011) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Peroxisome proliferator-activated receptor delta |
|---|
| Name: | Peroxisome proliferator-activated receptor delta |
|---|
| Synonyms: | NR1C2 | NUC1 | NUCI | Nuclear hormone receptor 1 | Nuclear receptor subfamily 1 group C member 2 | PPAR delta | PPAR-beta | PPARB | PPARD | PPARD_HUMAN | Peroxisome proliferator-activated receptor | Peroxisome proliferator-activated receptor beta | Peroxisome proliferator-activated receptor delta |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 49910.45 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | Q03181 |
|---|
| Residue: | 441 |
|---|
| Sequence: | MEQPQEEAPEVREEEEKEEVAEAEGAPELNGGPQHALPSSSYTDLSRSSSPPSLLDQLQM
GCDGASCGSLNMECRVCGDKASGFHYGVHACEGCKGFFRRTIRMKLEYEKCERSCKIQKK
NRNKCQYCRFQKCLALGMSHNAIRFGRMPEAEKRKLVAGLTANEGSQYNPQVADLKAFSK
HIYNAYLKNFNMTKKKARSILTGKASHTAPFVIHDIETLWQAEKGLVWKQLVNGLPPYKE
ISVHVFYRCQCTTVETVRELTEFAKSIPSFSSLFLNDQVTLLKYGVHEAIFAMLASIVNK
DGLLVANGSGFVTREFLRSLRKPFSDIIEPKFEFAVKFNALELDDSDLALFIAAIILCGD
RPGLMNVPRVEAIQDTILRALEFHLQANHPDAQYLFPKLLQKMADLRQLVTEHAQMMQRI
KKTETETSLHPLLQEIYKDMY
|
|
|
|---|
| BDBM28701 |
|---|
| n/a |
|---|
| Name | BDBM28701 |
|---|
| Synonyms: | 2-(4-{2-[(4-chlorophenyl)formamido]ethyl}phenoxy)-2-methylpropanoic acid | Azufibrat | Bezafibrate | CHEMBL264374 | Cedur | Sklerofibrat |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C19H20ClNO4 |
|---|
| Mol. Mass. | 361.819 |
|---|
| SMILES | CC(C)(Oc1ccc(CCNC(=O)c2ccc(Cl)cc2)cc1)C(O)=O |
|---|
| Structure | 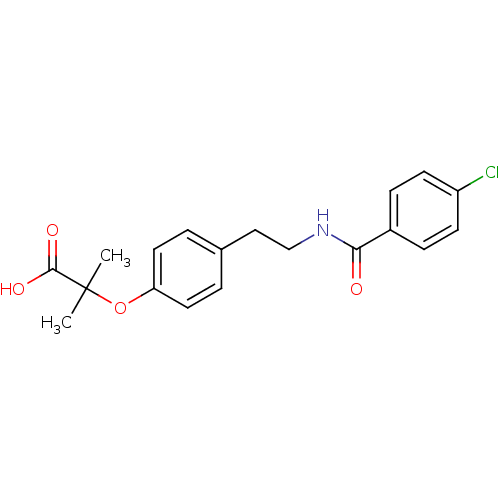
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Makadia, P; Shah, SR; Pingali, H; Zaware, P; Patel, D; Pola, S; Thube, B; Priyadarshini, P; Suthar, D; Shah, M; Giri, S; Trivedi, C; Jain, M; Patel, P; Bahekar, R Effect of structurally constrained oxime-ether linker on PPAR subtype selectivity: Discovery of a novel and potent series of PPAR-pan agonists. Bioorg Med Chem19:771-82 (2011) [PubMed] Article
Makadia, P; Shah, SR; Pingali, H; Zaware, P; Patel, D; Pola, S; Thube, B; Priyadarshini, P; Suthar, D; Shah, M; Giri, S; Trivedi, C; Jain, M; Patel, P; Bahekar, R Effect of structurally constrained oxime-ether linker on PPAR subtype selectivity: Discovery of a novel and potent series of PPAR-pan agonists. Bioorg Med Chem19:771-82 (2011) [PubMed] Article