| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Tyrosine-protein kinase ZAP-70 |
|---|
| Ligand | BDBM50346918 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_753067 (CHEMBL1798995) |
|---|
| IC50 | >10000±n/a nM |
|---|
| Citation |  Devadas, B; Selness, SR; Xing, L; Madsen, HM; Marrufo, LD; Shieh, H; Messing, DM; Yang, JZ; Morgan, HM; Anderson, GD; Webb, EG; Zhang, J; Devraj, RV; Monahan, JB Substituted N-aryl-6-pyrimidinones: a new class of potent, selective, and orally active p38 MAP kinase inhibitors. Bioorg Med Chem Lett21:3856-60 (2011) [PubMed] Article Devadas, B; Selness, SR; Xing, L; Madsen, HM; Marrufo, LD; Shieh, H; Messing, DM; Yang, JZ; Morgan, HM; Anderson, GD; Webb, EG; Zhang, J; Devraj, RV; Monahan, JB Substituted N-aryl-6-pyrimidinones: a new class of potent, selective, and orally active p38 MAP kinase inhibitors. Bioorg Med Chem Lett21:3856-60 (2011) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Tyrosine-protein kinase ZAP-70 |
|---|
| Name: | Tyrosine-protein kinase ZAP-70 |
|---|
| Synonyms: | 70 kDa zeta-associated protein | SRK | Syk-related tyrosine kinase | Tyrosine Kinase ZAP-70 | Tyrosine-protein kinase ZAP-70 | Tyrosine-protein kinase ZAP-70 (Syk) | Tyrosine-protein kinase ZAP-70 (ZAP70) | Tyrosine-protein kinase ZAP70 | ZAP70 | ZAP70_HUMAN | Zeta-chain (TCR) associated protein kinase 70kDa |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 69881.61 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | ZAP-70 SH2 domain was expressed and purified from E. coli using O-phospho-L-tyrosine-agarose column. |
|---|
| Residue: | 619 |
|---|
| Sequence: | MPDPAAHLPFFYGSISRAEAEEHLKLAGMADGLFLLRQCLRSLGGYVLSLVHDVRFHHFP
IERQLNGTYAIAGGKAHCGPAELCEFYSRDPDGLPCNLRKPCNRPSGLEPQPGVFDCLRD
AMVRDYVRQTWKLEGEALEQAIISQAPQVEKLIATTAHERMPWYHSSLTREEAERKLYSG
AQTDGKFLLRPRKEQGTYALSLIYGKTVYHYLISQDKAGKYCIPEGTKFDTLWQLVEYLK
LKADGLIYCLKEACPNSSASNASGAAAPTLPAHPSTLTHPQRRIDTLNSDGYTPEPARIT
SPDKPRPMPMDTSVYESPYSDPEELKDKKLFLKRDNLLIADIELGCGNFGSVRQGVYRMR
KKQIDVAIKVLKQGTEKADTEEMMREAQIMHQLDNPYIVRLIGVCQAEALMLVMEMAGGG
PLHKFLVGKREEIPVSNVAELLHQVSMGMKYLEEKNFVHRDLAARNVLLVNRHYAKISDF
GLSKALGADDSYYTARSAGKWPLKWYAPECINFRKFSSRSDVWSYGVTMWEALSYGQKPY
KKMKGPEVMAFIEQGKRMECPPECPPELYALMSDCWIYKWEDRPDFLTVEQRMRACYYSL
ASKVEGPPGSTQKAEAACA
|
|
|
|---|
| BDBM50346918 |
|---|
| n/a |
|---|
| Name | BDBM50346918 |
|---|
| Synonyms: | CHEMBL1738710 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C21H18ClF2N3O4 |
|---|
| Mol. Mass. | 449.835 |
|---|
| SMILES | Cc1ccc(cc1-n1cnc(OCc2ccc(F)cc2F)c(Cl)c1=O)C(=O)NCCO |
|---|
| Structure | 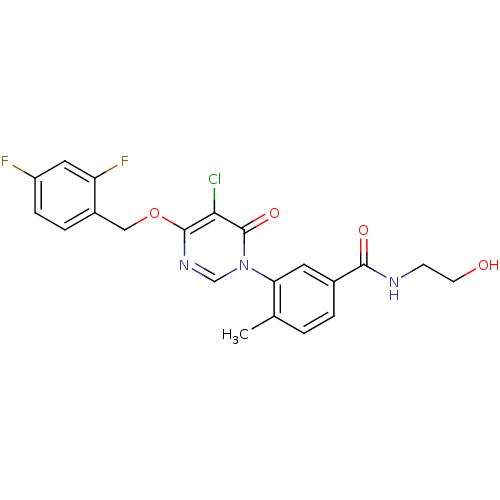
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Devadas, B; Selness, SR; Xing, L; Madsen, HM; Marrufo, LD; Shieh, H; Messing, DM; Yang, JZ; Morgan, HM; Anderson, GD; Webb, EG; Zhang, J; Devraj, RV; Monahan, JB Substituted N-aryl-6-pyrimidinones: a new class of potent, selective, and orally active p38 MAP kinase inhibitors. Bioorg Med Chem Lett21:3856-60 (2011) [PubMed] Article
Devadas, B; Selness, SR; Xing, L; Madsen, HM; Marrufo, LD; Shieh, H; Messing, DM; Yang, JZ; Morgan, HM; Anderson, GD; Webb, EG; Zhang, J; Devraj, RV; Monahan, JB Substituted N-aryl-6-pyrimidinones: a new class of potent, selective, and orally active p38 MAP kinase inhibitors. Bioorg Med Chem Lett21:3856-60 (2011) [PubMed] Article