| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Tyrosine-protein kinase ZAP-70 |
|---|
| Ligand | BDBM50128285 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_774224 (CHEMBL1908441) |
|---|
| Kd | >10000±n/a nM |
|---|
| Citation |  Davis, MI; Hunt, JP; Herrgard, S; Ciceri, P; Wodicka, LM; Pallares, G; Hocker, M; Treiber, DK; Zarrinkar, PP Comprehensive analysis of kinase inhibitor selectivity. Nat Biotechnol29:1046-51 (2011) [PubMed] Article Davis, MI; Hunt, JP; Herrgard, S; Ciceri, P; Wodicka, LM; Pallares, G; Hocker, M; Treiber, DK; Zarrinkar, PP Comprehensive analysis of kinase inhibitor selectivity. Nat Biotechnol29:1046-51 (2011) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Tyrosine-protein kinase ZAP-70 |
|---|
| Name: | Tyrosine-protein kinase ZAP-70 |
|---|
| Synonyms: | 70 kDa zeta-associated protein | SRK | Syk-related tyrosine kinase | Tyrosine Kinase ZAP-70 | Tyrosine-protein kinase ZAP-70 | Tyrosine-protein kinase ZAP-70 (Syk) | Tyrosine-protein kinase ZAP-70 (ZAP70) | Tyrosine-protein kinase ZAP70 | ZAP70 | ZAP70_HUMAN | Zeta-chain (TCR) associated protein kinase 70kDa |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 69881.61 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | ZAP-70 SH2 domain was expressed and purified from E. coli using O-phospho-L-tyrosine-agarose column. |
|---|
| Residue: | 619 |
|---|
| Sequence: | MPDPAAHLPFFYGSISRAEAEEHLKLAGMADGLFLLRQCLRSLGGYVLSLVHDVRFHHFP
IERQLNGTYAIAGGKAHCGPAELCEFYSRDPDGLPCNLRKPCNRPSGLEPQPGVFDCLRD
AMVRDYVRQTWKLEGEALEQAIISQAPQVEKLIATTAHERMPWYHSSLTREEAERKLYSG
AQTDGKFLLRPRKEQGTYALSLIYGKTVYHYLISQDKAGKYCIPEGTKFDTLWQLVEYLK
LKADGLIYCLKEACPNSSASNASGAAAPTLPAHPSTLTHPQRRIDTLNSDGYTPEPARIT
SPDKPRPMPMDTSVYESPYSDPEELKDKKLFLKRDNLLIADIELGCGNFGSVRQGVYRMR
KKQIDVAIKVLKQGTEKADTEEMMREAQIMHQLDNPYIVRLIGVCQAEALMLVMEMAGGG
PLHKFLVGKREEIPVSNVAELLHQVSMGMKYLEEKNFVHRDLAARNVLLVNRHYAKISDF
GLSKALGADDSYYTARSAGKWPLKWYAPECINFRKFSSRSDVWSYGVTMWEALSYGQKPY
KKMKGPEVMAFIEQGKRMECPPECPPELYALMSDCWIYKWEDRPDFLTVEQRMRACYYSL
ASKVEGPPGSTQKAEAACA
|
|
|
|---|
| BDBM50128285 |
|---|
| n/a |
|---|
| Name | BDBM50128285 |
|---|
| Synonyms: | 3-(1-Methyl-1H-indol-3-yl)-4-[1-(1-pyridin-2-ylmethyl-piperidin-4-yl)-1H-indol-3-yl]-pyrrole-2,5-dione | CHEMBL300138 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C32H29N5O2 |
|---|
| Mol. Mass. | 515.605 |
|---|
| SMILES | Cn1cc(C2=C(C(=O)NC2=O)c2cn(C3CCN(Cc4ccccn4)CC3)c3ccccc23)c2ccccc12 |t:4| |
|---|
| Structure | 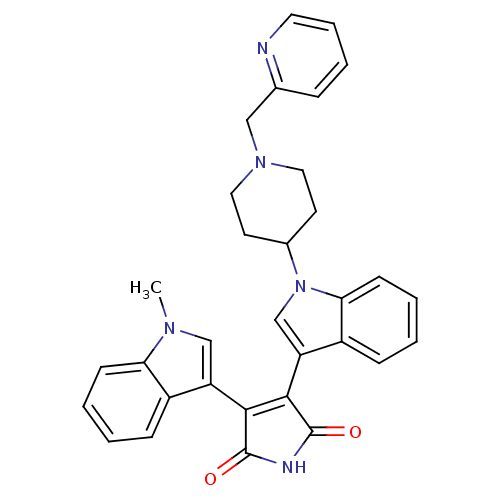
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Davis, MI; Hunt, JP; Herrgard, S; Ciceri, P; Wodicka, LM; Pallares, G; Hocker, M; Treiber, DK; Zarrinkar, PP Comprehensive analysis of kinase inhibitor selectivity. Nat Biotechnol29:1046-51 (2011) [PubMed] Article
Davis, MI; Hunt, JP; Herrgard, S; Ciceri, P; Wodicka, LM; Pallares, G; Hocker, M; Treiber, DK; Zarrinkar, PP Comprehensive analysis of kinase inhibitor selectivity. Nat Biotechnol29:1046-51 (2011) [PubMed] Article