

 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data
| Reaction Details | |||
|---|---|---|---|
 | Report a problem with these data | ||
| Target | Dipeptidyl peptidase 8 | ||
| Ligand | BDBM50356589 | ||
| Substrate/Competitor | n/a | ||
| Meas. Tech. | ChEMBL_776112 (CHEMBL1912808) | ||
| Ki | 3477±n/a nM | ||
| Citation |  Wang, W; Devasthale, P; Wang, A; Harrity, T; Egan, D; Morgan, N; Cap, M; Fura, A; Klei, HE; Kish, K; Weigelt, C; Sun, L; Levesque, P; Li, YX; Zahler, R; Kirby, MS; Hamann, LG 7-Oxopyrrolopyridine-derived DPP4 inhibitors-mitigation of CYP and hERG liabilities via introduction of polar functionalities in the active site. Bioorg Med Chem Lett21:6646-51 (2011) [PubMed] Article Wang, W; Devasthale, P; Wang, A; Harrity, T; Egan, D; Morgan, N; Cap, M; Fura, A; Klei, HE; Kish, K; Weigelt, C; Sun, L; Levesque, P; Li, YX; Zahler, R; Kirby, MS; Hamann, LG 7-Oxopyrrolopyridine-derived DPP4 inhibitors-mitigation of CYP and hERG liabilities via introduction of polar functionalities in the active site. Bioorg Med Chem Lett21:6646-51 (2011) [PubMed] Article | ||
| More Info.: | Get all data from this article, Assay Method | ||
| Dipeptidyl peptidase 8 | |||
| Name: | Dipeptidyl peptidase 8 | ||
| Synonyms: | DPP8 | DPP8_HUMAN | DPRP-1 | DPRP1 | Dipeptidyl peptidase 8 (DPP-8) | Dipeptidyl peptidase 8 (DPP8) | Dipeptidyl peptidase 8/9 | Dipeptidyl peptidase IV-related protein 1 | Dipeptidyl peptidase VIII | Dipeptidyl peptidase VIII (DDP-VIII) | Prolyl dipeptidase DPP8 | ||
| Type: | Enzyme | ||
| Mol. Mass.: | 103342.62 | ||
| Organism: | Homo sapiens (Human) | ||
| Description: | Q6V1X1 | ||
| Residue: | 898 | ||
| Sequence: |
| ||
| BDBM50356589 | |||
| n/a | |||
| Name | BDBM50356589 | ||
| Synonyms: | CHEMBL1910119 | ||
| Type | Small organic molecule | ||
| Emp. Form. | C23H25Cl2N5O3 | ||
| Mol. Mass. | 490.382 | ||
| SMILES | Cc1nc2C(=O)N(CC(=O)N3CCC(CC3)C(N)=O)Cc2c(c1CN)-c1ccc(Cl)cc1Cl |(6.41,-30.16,;5.08,-30.93,;3.75,-30.17,;2.42,-30.95,;.95,-30.47,;.48,-29,;.05,-31.71,;-1.49,-31.71,;-2.25,-33.05,;-1.48,-34.38,;-3.79,-33.05,;-4.56,-31.72,;-6.09,-31.72,;-6.87,-33.05,;-6.1,-34.38,;-4.56,-34.39,;-8.41,-33.04,;-9.18,-31.71,;-9.18,-34.38,;.95,-32.96,;2.41,-32.48,;3.75,-33.26,;5.09,-32.48,;6.42,-33.25,;7.75,-32.48,;3.75,-34.79,;2.41,-35.56,;2.41,-37.1,;3.75,-37.87,;3.75,-39.41,;5.09,-37.09,;5.08,-35.56,;6.41,-34.78,)| | ||
| Structure | 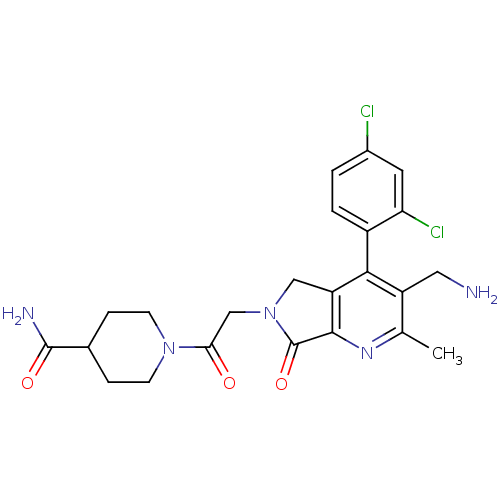
| ||