| Reaction Details |
|---|
 | Report a problem with these data |
| Target | 5-hydroxytryptamine receptor 1A |
|---|
| Ligand | BDBM50063260 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_1304 (CHEMBL616681) |
|---|
| Ki | 24000±n/a nM |
|---|
| Citation |  Cappelli, A; Anzini, M; Vomero, S; Mennuni, L; Makovec, F; Doucet, E; Hamon, M; Bruni, G; Romeo, MR; Menziani, MC; De Benedetti, PG; Langer, T Novel potent and selective central 5-HT3 receptor ligands provided with different intrinsic efficacy. 1. Mapping the central 5-HT3 receptor binding site by arylpiperazine derivatives. J Med Chem41:728-41 (1998) [PubMed] Article Cappelli, A; Anzini, M; Vomero, S; Mennuni, L; Makovec, F; Doucet, E; Hamon, M; Bruni, G; Romeo, MR; Menziani, MC; De Benedetti, PG; Langer, T Novel potent and selective central 5-HT3 receptor ligands provided with different intrinsic efficacy. 1. Mapping the central 5-HT3 receptor binding site by arylpiperazine derivatives. J Med Chem41:728-41 (1998) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| 5-hydroxytryptamine receptor 1A |
|---|
| Name: | 5-hydroxytryptamine receptor 1A |
|---|
| Synonyms: | 5-HT-1A | 5-HT1 | 5-HT1A | 5-Hydroxytryptamine receptor 1A (5-HT1A) | 5-hydroxytryptamine receptor 1A (5HT1A) | 5HT1A_RAT | 5ht1a | G-21 | Htr1a | Serotonin 1 (5-HT1) receptor | Serotonin 1a (5-HT1a) receptor/Adrenergic receptor alpha-1 | Serotonin receptor 1A |
|---|
| Type: | G Protein-Coupled Receptor (GPCR) |
|---|
| Mol. Mass.: | 46445.29 |
|---|
| Organism: | Rattus norvegicus (rat) |
|---|
| Description: | Binding assays were performed using rat hippocampal membranes. |
|---|
| Residue: | 422 |
|---|
| Sequence: | MDVFSFGQGNNTTASQEPFGTGGNVTSISDVTFSYQVITSLLLGTLIFCAVLGNACVVAA
IALERSLQNVANYLIGSLAVTDLMVSVLVLPMAALYQVLNKWTLGQVTCDLFIALDVLCC
TSSILHLCAIALDRYWAITDPIDYVNKRTPRRAAALISLTWLIGFLISIPPMLGWRTPED
RSDPDACTISKDHGYTIYSTFGAFYIPLLLMLVLYGRIFRAARFRIRKTVRKVEKKGAGT
SLGTSSAPPPKKSLNGQPGSGDWRRCAENRAVGTPCTNGAVRQGDDEATLEVIEVHRVGN
SKEHLPLPSESGSNSYAPACLERKNERNAEAKRKMALARERKTVKTLGIIMGTFILCWLP
FFIVALVLPFCESSCHMPALLGAIINWLGYSNSLLNPVIYAYFNKDFQNAFKKIIKCKFC
RR
|
|
|
|---|
| BDBM50063260 |
|---|
| n/a |
|---|
| Name | BDBM50063260 |
|---|
| Synonyms: | 10-(4-methylhexahydro-1-pyrazinyl)-9-azatetracyclo[11.2.1.02,11.03,8]hexadeca-2(11),3,5,7,9-pentaene | 6-(4-Methyl-piperazin-1-yl)-7,8,9,10-tetrahydro-8,10-ethanophenanthridine | CHEMBL287987 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C20H25N3 |
|---|
| Mol. Mass. | 307.4326 |
|---|
| SMILES | CN1CCN(CC1)c1nc2ccccc2c2C3CCC(C3)Cc12 |
|---|
| Structure | 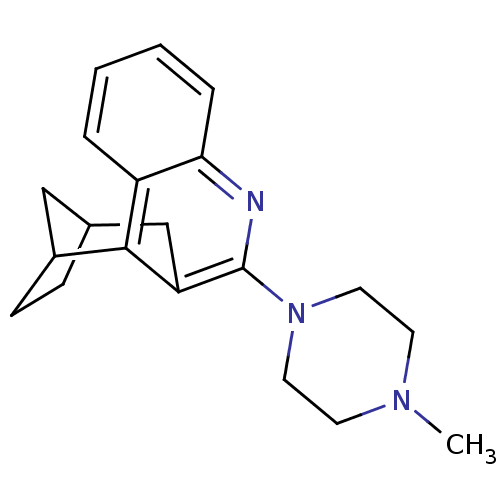
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Cappelli, A; Anzini, M; Vomero, S; Mennuni, L; Makovec, F; Doucet, E; Hamon, M; Bruni, G; Romeo, MR; Menziani, MC; De Benedetti, PG; Langer, T Novel potent and selective central 5-HT3 receptor ligands provided with different intrinsic efficacy. 1. Mapping the central 5-HT3 receptor binding site by arylpiperazine derivatives. J Med Chem41:728-41 (1998) [PubMed] Article
Cappelli, A; Anzini, M; Vomero, S; Mennuni, L; Makovec, F; Doucet, E; Hamon, M; Bruni, G; Romeo, MR; Menziani, MC; De Benedetti, PG; Langer, T Novel potent and selective central 5-HT3 receptor ligands provided with different intrinsic efficacy. 1. Mapping the central 5-HT3 receptor binding site by arylpiperazine derivatives. J Med Chem41:728-41 (1998) [PubMed] Article