| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Sucrase-isomaltase, intestinal |
|---|
| Ligand | BDBM50031481 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_472704 (CHEMBL923888) |
|---|
| IC50 | 290000±n/a nM |
|---|
| Citation |  Minami, Y; Kuriyama, C; Ikeda, K; Kato, A; Takebayashi, K; Adachi, I; Fleet, GW; Kettawan, A; Okamoto, T; Asano, N Effect of five-membered sugar mimics on mammalian glycogen-degrading enzymes and various glucosidases. Bioorg Med Chem16:2734-40 (2008) [PubMed] Article Minami, Y; Kuriyama, C; Ikeda, K; Kato, A; Takebayashi, K; Adachi, I; Fleet, GW; Kettawan, A; Okamoto, T; Asano, N Effect of five-membered sugar mimics on mammalian glycogen-degrading enzymes and various glucosidases. Bioorg Med Chem16:2734-40 (2008) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Sucrase-isomaltase, intestinal |
|---|
| Name: | Sucrase-isomaltase, intestinal |
|---|
| Synonyms: | SUIS_RAT | Si | Sucrase-isomaltase | alpha-Glucosidase (α-Glucosidase) |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 210329.04 |
|---|
| Organism: | Rattus norvegicus (Rat) |
|---|
| Description: | P23739 |
|---|
| Residue: | 1841 |
|---|
| Sequence: | MAKKKFSALEISLIVLFIIVTAIAIALVTVLATKVPAVEEIKSPTPTSNSTPTSTPTSTS
TPTSTSTPSPGKCPPEQGEPINERINCIPEQHPTKAICEERGCCWRPWNNTVIPWCFFAD
NHGYNAESITNENAGLKATLNRIPSPTLFGEDIKSVILTTQTQTGNRFRFKITDPNNKRY
EVPHQFVKEETGIPAADTLYDVQVSENPFSIKVIRKSNNKVLCDTSVGPLLYSNQYLQIS
TRLPSEYIYGFGGHIHKRFRHDLYWKTWPIFTRDEIPGDNNHNLYGHQTFFMGIGDTSGK
SYGVFLMNSNAMEVFIQPTPIITYRVTGGILDFYIFLGDTPEQVVQQYQEVHWRPAMPAY
WNLGFQLSRWNYGSLDTVSEVVRRNREAGIPYDAQVTDIDYMEDHKEFTYDRVKFNGLPE
FAQDLHNHGKYIIILDPAISINKRANGAEYQTYVRGNEKNVWVNESDGTTPLIGEVWPGL
TVYPDFTNPQTIEWWANECNLFHQQVEYDGLWIDMNEVSSFIQGSLNLKGVLLIVLNYPP
FTPGILDKVMYSKTLCMDAVQHWGKQYDVHSLYGYSMAIATEQAVERVFPNKRSFILTRS
TFGGSGRHANHWLGDNTASWEQMEWSITGMLEFGIFGMPLVGATSCGFLADTTEELCRRW
MQLGAFYPFSRNHNAEGYMEQDPAYFGQDSSRHYLTIRYTLLPFLYTLFYRAHMFGETVA
RPFLYEFYDDTNSWIEDTQFLWGPALLITPVLRPGVENVSAYIPNATWYDYETGIKRPWR
KERINMYLPGDKIGLHLRGGYIIPTQEPDVTTTASRKNPLGLIVALDDNQAAKGELFWDD
GESKDSIEKKMYILYTFSVSNNELVLNCTHSSYAEGTSLAFKTIKVLGLREDVRSITVGE
NDQQMATHTNFTFDSANKILSITALNFNLAGSFIVRWCRTFSDNEKFTCYPDVGTATEGT
CTQRGCLWQPVSGLSNVPPYYFPPENNPYTLTSIQPLPTGITAELQLNPPNARIKLPSNP
ISTLRVGVKYHPNDMLQFKIYDAQHKRYEVPVPLNIPDTPTSSNERLYDVEIKENPFGIQ
VRRRSSGKLIWDSRLPGFGFNDQFIQISTRLPSNYLYGFGEVEHTAFKRDLNWHTWGMFT
RDQPPGYKLNSYGFHPYYMALENEGNAHGVLLLNSNGMDVTFQPTPALTYRTIGGILDFY
MFLGPTPEIATRQYHEVIGFPVMPPYWALGFQLCRYGYRNTSEIEQLYNDMVAANIPYDV
QYTDINYMERQLDFTIGERFKTLPEFVDRIRKDGMKYIVILAPAISGNETQPYPAFERGI
QKDVFVKWPNTNDICWPKVWPDLPNVTIDETITEDEAVNASRAHVAFPDFFRNSTLEWWA
REIYDFYNEKMKFDGLWIDMNEPSSFGIQMGGKVLNECRRMMTLNYPPVFSPELRVKEGE
GASISEAMCMETEHILIDGSSVLQYDVHNLYGWSQVKPTLDALQNTTGLRGIVISRSTYP
TTGRWGGHWLGDNYTTWDNLEKSLIGMLELNLFGIPYIGADICGVFHDSGYPSLYFVGIQ
VGAFYPYPRESPTINFTRSQDPVSWMKLLLQMSKKVLEIRYTLLPYFYTQMHEAHAHGGT
VIRPLMHEFFDDKETWEIYKQFLWGPAFMVTPVVEPFRTSVTGYVPKARWFDYHTGADIK
LKGILHTFSAPFDTINLHVRGGYILPCQEPARNTHLSRQNYMKLIVAADDNQMAQGTLFG
DDGESIDTYERGQYTSIQFNLNQTTLTSTVLANGYKNKQEMRLGSIHIWGKGTLRISNAN
LVYGGRKHQPPFTQEEAKETLIFDLKNMNVTLDEPIQITWS
|
|
|
|---|
| BDBM50031481 |
|---|
| n/a |
|---|
| Name | BDBM50031481 |
|---|
| Synonyms: | (2R,3R,4R,5R)-2,5-Bis-hydroxymethyl-pyrrolidine-3,4-diol | 2(R),5(R)-bis(hydroxymethyl)-3(R),4(R)-dihydroxypyrrolidine | 2R,5R-dihydroxymethyl-3R,4R-dihydroxypyrrolidine | CHEMBL312653 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C6H13NO4 |
|---|
| Mol. Mass. | 163.1717 |
|---|
| SMILES | OC[C@H]1N[C@H](CO)[C@@H](O)[C@@H]1O |r| |
|---|
| Structure | 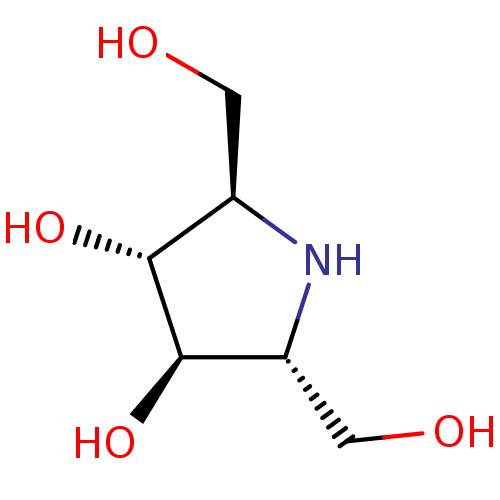
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Minami, Y; Kuriyama, C; Ikeda, K; Kato, A; Takebayashi, K; Adachi, I; Fleet, GW; Kettawan, A; Okamoto, T; Asano, N Effect of five-membered sugar mimics on mammalian glycogen-degrading enzymes and various glucosidases. Bioorg Med Chem16:2734-40 (2008) [PubMed] Article
Minami, Y; Kuriyama, C; Ikeda, K; Kato, A; Takebayashi, K; Adachi, I; Fleet, GW; Kettawan, A; Okamoto, T; Asano, N Effect of five-membered sugar mimics on mammalian glycogen-degrading enzymes and various glucosidases. Bioorg Med Chem16:2734-40 (2008) [PubMed] Article