

 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data
| Reaction Details | |||
|---|---|---|---|
 | Report a problem with these data | ||
| Target | Diacylglycerol O-acyltransferase 1 | ||
| Ligand | BDBM50117187 | ||
| Substrate/Competitor | n/a | ||
| Meas. Tech. | ChEMBL_1513968 (CHEMBL3616154) | ||
| IC50 | 5.0±n/a nM | ||
| Citation |  Pagire, SH; Pagire, HS; Lee, GB; Han, SJ; Kwak, HJ; Kim, JY; Kim, KY; Rhee, SD; Ryu, JI; Song, JS; Bae, MA; Park, MJ; Kim, D; Lee, DH; Ahn, JH Discovery and optimization of adamantane carboxylic acid derivatives as potent diacylglycerol acyltransferase 1 inhibitors for the potential treatment of obesity and diabetes. Eur J Med Chem101:716-35 (2015) [PubMed] Article Pagire, SH; Pagire, HS; Lee, GB; Han, SJ; Kwak, HJ; Kim, JY; Kim, KY; Rhee, SD; Ryu, JI; Song, JS; Bae, MA; Park, MJ; Kim, D; Lee, DH; Ahn, JH Discovery and optimization of adamantane carboxylic acid derivatives as potent diacylglycerol acyltransferase 1 inhibitors for the potential treatment of obesity and diabetes. Eur J Med Chem101:716-35 (2015) [PubMed] Article | ||
| More Info.: | Get all data from this article, Assay Method | ||
| Diacylglycerol O-acyltransferase 1 | |||
| Name: | Diacylglycerol O-acyltransferase 1 | ||
| Synonyms: | DGAT1_MOUSE | Dgat | Dgat1 | Diacyl Glycerolacyltransferase 1 (DGAT-1) | Diacylglycerol O-acyltransferase 1 (DGAT1) | Diglyceride acyltransferase | ||
| Type: | Enzyme | ||
| Mol. Mass.: | 56810.61 | ||
| Organism: | Mus musculus (mouse) | ||
| Description: | In this assay, recombinant mouse DGAT-1 containing an N-terminal His6-epitope tag was produced in the baculovirus expression system. | ||
| Residue: | 498 | ||
| Sequence: |
| ||
| BDBM50117187 | |||
| n/a | |||
| Name | BDBM50117187 | ||
| Synonyms: | CHEMBL3613341 | ||
| Type | Small organic molecule | ||
| Emp. Form. | C31H30F3N3O6 | ||
| Mol. Mass. | 597.5816 | ||
| SMILES | OC(=O)C12CC3CC(C1)C(Oc1ccc(cc1)C(=O)NCCNC(=O)c1nc(oc1C(F)(F)F)-c1ccccc1)C(C3)C2 |TLB:6:5:42:8.7.9,6:7:4.5.41:42,THB:9:7:4:41.40.42,9:40:4:8.6.7,10:9:4.5.41:42,(3.79,.17,;3.07,1.17,;3.58,2.29,;1.56,1.02,;1.56,2.59,;.12,3.07,;-1.2,2.69,;-1.2,1.02,;.34,.44,;-2.19,-.22,;-3.39,-1.16,;-3.17,-2.69,;-4.38,-3.64,;-4.16,-5.17,;-2.73,-5.74,;-1.52,-4.78,;-1.74,-3.26,;-2.5,-7.26,;-3.47,-8.03,;-1.07,-7.83,;-.85,-9.36,;.59,-9.92,;.81,-11.45,;2.24,-12.02,;3.21,-11.25,;2.47,-13.54,;1.36,-14.59,;2.04,-15.98,;3.57,-15.75,;3.83,-14.24,;5.2,-13.55,;5.27,-12.32,;6.23,-14.22,;6.3,-12.99,;1.32,-17.34,;2.04,-18.7,;1.23,-20,;-.31,-19.94,;-1.03,-18.58,;-.22,-17.28,;-.95,.32,;-1,2.05,;.56,-.22,)| | ||
| Structure | 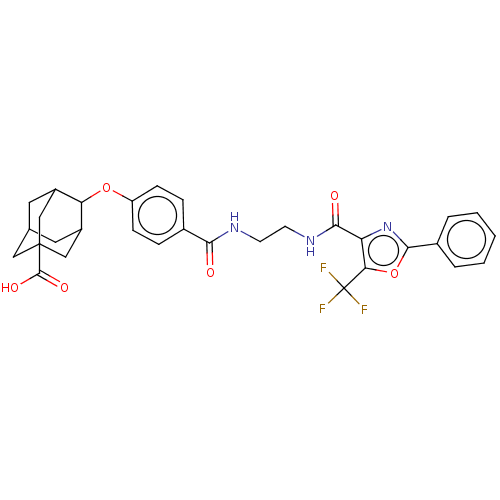
| ||