| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Serine/threonine-protein kinase TBK1 |
|---|
| Ligand | BDBM2621 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | KinEASE-STK Assay |
|---|
| IC50 | 1.66e+3± 709 nM |
|---|
| Citation |  Richters, A; Basu, D; Engel, J; Ercanoglu, MS; Balke-Want, H; Tesch, R; Thomas, RK; Rauh, D Identification and further development of potent TBK1 inhibitors. ACS Chem Biol10:289-98 (2015) [PubMed] Article Richters, A; Basu, D; Engel, J; Ercanoglu, MS; Balke-Want, H; Tesch, R; Thomas, RK; Rauh, D Identification and further development of potent TBK1 inhibitors. ACS Chem Biol10:289-98 (2015) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Serine/threonine-protein kinase TBK1 |
|---|
| Name: | Serine/threonine-protein kinase TBK1 |
|---|
| Synonyms: | NAK | NF-kappa-B-activating kinase | Protein cereblon/Serine/threonine-protein kinase TBK1 | T2K | TANK-binding kinase 1 (TBK-1) | TANK-binding kinase 1 (TBK1) | TBK1 | TBK1_HUMAN |
|---|
| Type: | protein |
|---|
| Mol. Mass.: | 83645.20 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | Q9UHD2 |
|---|
| Residue: | 729 |
|---|
| Sequence: | MQSTSNHLWLLSDILGQGATANVFRGRHKKTGDLFAIKVFNNISFLRPVDVQMREFEVLK
KLNHKNIVKLFAIEEETTTRHKVLIMEFCPCGSLYTVLEEPSNAYGLPESEFLIVLRDVV
GGMNHLRENGIVHRDIKPGNIMRVIGEDGQSVYKLTDFGAARELEDDEQFVSLYGTEEYL
HPDMYERAVLRKDHQKKYGATVDLWSIGVTFYHAATGSLPFRPFEGPRRNKEVMYKIITG
KPSGAISGVQKAENGPIDWSGDMPVSCSLSRGLQVLLTPVLANILEADQEKCWGFDQFFA
ETSDILHRMVIHVFSLQQMTAHKIYIHSYNTATIFHELVYKQTKIISSNQELIYEGRRLV
LEPGRLAQHFPKTTEENPIFVVSREPLNTIGLIYEKISLPKVHPRYDLDGDASMAKAITG
VVCYACRIASTLLLYQELMRKGIRWLIELIKDDYNETVHKKTEVVITLDFCIRNIEKTVK
VYEKLMKINLEAAELGEISDIHTKLLRLSSSQGTIETSLQDIDSRLSPGGSLADAWAHQE
GTHPKDRNVEKLQVLLNCMTEIYYQFKKDKAERRLAYNEEQIHKFDKQKLYYHATKAMTH
FTDECVKKYEAFLNKSEEWIRKMLHLRKQLLSLTNQCFDIEEEVSKYQEYTNELQETLPQ
KMFTASSGIKHTMTPIYPSSNTLVEMTLGMKKLKEEMEGVVKELAENNHILERFGSLTMD
GGLRNVDCL
|
|
|
|---|
| BDBM2621 |
|---|
| n/a |
|---|
| Name | BDBM2621 |
|---|
| Synonyms: | 3-(6-methoxy-1-methyl-1H-indol-3-yl)-4-(1-methyl-1H-indol-3-yl)-2,5-dihydro-1H-pyrrole-2,5-dione | Bisindolylmaleimide deriv. 43 | ROCHE screening, 45 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C23H19N3O3 |
|---|
| Mol. Mass. | 385.4153 |
|---|
| SMILES | COc1ccc2c(cn(C)c2c1)C1=C(C(=O)NC1=O)c1cn(C)c2ccccc12 |t:14| |
|---|
| Structure | 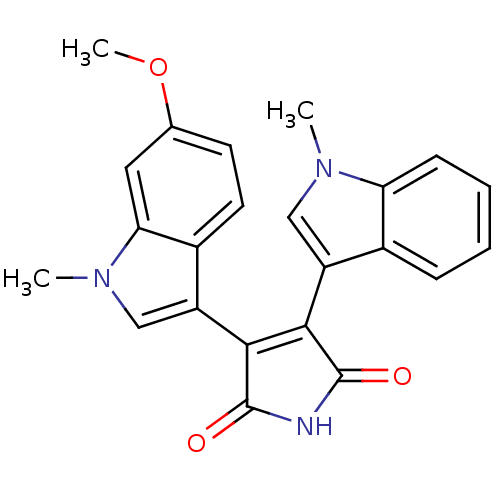
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Richters, A; Basu, D; Engel, J; Ercanoglu, MS; Balke-Want, H; Tesch, R; Thomas, RK; Rauh, D Identification and further development of potent TBK1 inhibitors. ACS Chem Biol10:289-98 (2015) [PubMed] Article
Richters, A; Basu, D; Engel, J; Ercanoglu, MS; Balke-Want, H; Tesch, R; Thomas, RK; Rauh, D Identification and further development of potent TBK1 inhibitors. ACS Chem Biol10:289-98 (2015) [PubMed] Article