

 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data
| Target/Host (Institution) | Ligand | Target/Host Links | Ligand Links | Trg + Lig Links | Ki nM | ΔG° kJ/mole | IC50 nM | Kd nM | EC50/IC50 nM | koff s-1 | kon M-1s-1 | pH | Temp °C |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Ubiquitin carboxyl-terminal hydrolase 2 (Homo sapiens (Human)) | BDBM50458422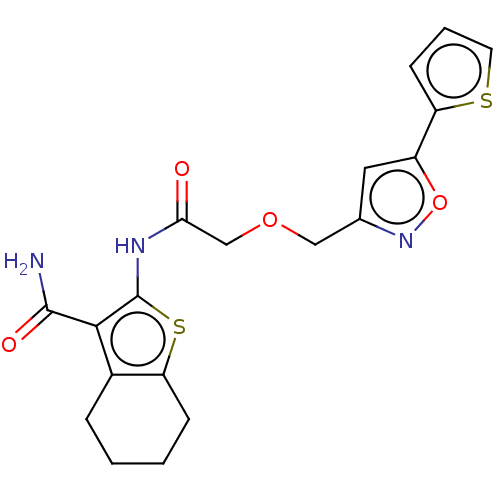 (CHEMBL4216958) | PDB MMDB KEGG UniProtKB/SwissProt B.MOAD antibodypedia GoogleScholar AffyNet | Purchase MCE PC cid PC sid UniChem | Article PubMed | n/a | n/a | 3.30E+3 | n/a | n/a | n/a | n/a | n/a | n/a |
Jagiellonian University Curated by ChEMBL | Assay Description Inhibition of human USP2a (258 to 605 residues) expressed in Escherichia coli BL21(DE3) using Ub-AMC as substrate incubated for 30 mins measured at 3... | Eur J Med Chem 150: 261-267 (2018) Article DOI: 10.1016/j.ejmech.2018.03.009 BindingDB Entry DOI: 10.7270/Q28G8P98 | |||||||||||
| More data for this Ligand-Target Pair | |||||||||||||
| Ubiquitin carboxyl-terminal hydrolase 2 (Homo sapiens (Human)) | BDBM50458415 (CHEMBL567508 | EN300-02284) | PDB MMDB KEGG UniProtKB/SwissProt B.MOAD antibodypedia GoogleScholar AffyNet | Purchase MCE PC cid PC sid UniChem | Article PubMed | n/a | n/a | 8.70E+3 | n/a | n/a | n/a | n/a | n/a | n/a |
Jagiellonian University Curated by ChEMBL | Assay Description Inhibition of human USP2a (258 to 605 residues) expressed in Escherichia coli BL21(DE3) using Ub-AMC as substrate incubated for 30 mins measured at 3... | Eur J Med Chem 150: 261-267 (2018) Article DOI: 10.1016/j.ejmech.2018.03.009 BindingDB Entry DOI: 10.7270/Q28G8P98 | |||||||||||
| More data for this Ligand-Target Pair | |||||||||||||
| Ubiquitin carboxyl-terminal hydrolase 2 (Homo sapiens (Human)) | BDBM50458419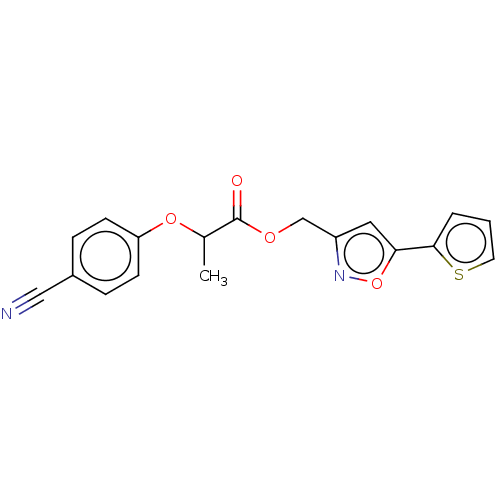 (CHEMBL4203286) | PDB MMDB KEGG UniProtKB/SwissProt B.MOAD antibodypedia GoogleScholar AffyNet | Purchase PC cid PC sid UniChem | Article PubMed | n/a | n/a | >5.00E+4 | n/a | n/a | n/a | n/a | n/a | n/a |
Jagiellonian University Curated by ChEMBL | Assay Description Inhibition of human USP2a (258 to 605 residues) expressed in Escherichia coli BL21(DE3) using Ub-AMC as substrate incubated for 30 mins measured at 3... | Eur J Med Chem 150: 261-267 (2018) Article DOI: 10.1016/j.ejmech.2018.03.009 BindingDB Entry DOI: 10.7270/Q28G8P98 | |||||||||||
| More data for this Ligand-Target Pair | |||||||||||||
| Ubiquitin carboxyl-terminal hydrolase 2 (Homo sapiens (Human)) | BDBM50458417 (CHEMBL4203489) | PDB MMDB KEGG UniProtKB/SwissProt B.MOAD antibodypedia GoogleScholar AffyNet | Purchase PC cid PC sid UniChem | Article PubMed | n/a | n/a | >1.00E+5 | n/a | n/a | n/a | n/a | n/a | n/a |
Jagiellonian University Curated by ChEMBL | Assay Description Inhibition of human USP2a (258 to 605 residues) expressed in Escherichia coli BL21(DE3) using Ub-AMC as substrate incubated for 30 mins measured at 3... | Eur J Med Chem 150: 261-267 (2018) Article DOI: 10.1016/j.ejmech.2018.03.009 BindingDB Entry DOI: 10.7270/Q28G8P98 | |||||||||||
| More data for this Ligand-Target Pair | |||||||||||||
| Ubiquitin carboxyl-terminal hydrolase 2 (Homo sapiens (Human)) | BDBM50458416 (CHEMBL1230155) | PDB MMDB KEGG UniProtKB/SwissProt B.MOAD antibodypedia GoogleScholar AffyNet | Purchase MCE PC cid PC sid PDB UniChem | Article PubMed | n/a | n/a | 1.30E+6 | n/a | n/a | n/a | n/a | n/a | n/a |
Jagiellonian University Curated by ChEMBL | Assay Description Inhibition of human USP2a (258 to 605 residues) expressed in Escherichia coli BL21(DE3) using Ub-AMC as substrate incubated for 30 mins measured at 3... | Eur J Med Chem 150: 261-267 (2018) Article DOI: 10.1016/j.ejmech.2018.03.009 BindingDB Entry DOI: 10.7270/Q28G8P98 | |||||||||||
| More data for this Ligand-Target Pair | |||||||||||||
| Ubiquitin carboxyl-terminal hydrolase 2 (Homo sapiens (Human)) | BDBM50458421 (CHEMBL1650269) | PDB MMDB KEGG UniProtKB/SwissProt B.MOAD antibodypedia GoogleScholar AffyNet | Purchase PC cid PC sid UniChem | Article PubMed | n/a | n/a | >5.00E+6 | n/a | n/a | n/a | n/a | n/a | n/a |
Jagiellonian University Curated by ChEMBL | Assay Description Inhibition of human USP2a (258 to 605 residues) expressed in Escherichia coli BL21(DE3) using Ub-AMC as substrate incubated for 30 mins measured at 3... | Eur J Med Chem 150: 261-267 (2018) Article DOI: 10.1016/j.ejmech.2018.03.009 BindingDB Entry DOI: 10.7270/Q28G8P98 | |||||||||||
| More data for this Ligand-Target Pair | |||||||||||||
| Ubiquitin carboxyl-terminal hydrolase 2 (Homo sapiens (Human)) | BDBM50458420 (CHEMBL4207620) | PDB MMDB KEGG UniProtKB/SwissProt B.MOAD antibodypedia GoogleScholar AffyNet | Purchase MCE PC cid PC sid UniChem | Article PubMed | n/a | n/a | >5.00E+6 | n/a | n/a | n/a | n/a | n/a | n/a |
Jagiellonian University Curated by ChEMBL | Assay Description Inhibition of human USP2a (258 to 605 residues) expressed in Escherichia coli BL21(DE3) using Ub-AMC as substrate incubated for 30 mins measured at 3... | Eur J Med Chem 150: 261-267 (2018) Article DOI: 10.1016/j.ejmech.2018.03.009 BindingDB Entry DOI: 10.7270/Q28G8P98 | |||||||||||
| More data for this Ligand-Target Pair | |||||||||||||
| Ubiquitin carboxyl-terminal hydrolase 2 (Homo sapiens (Human)) | BDBM50458418 (CHEMBL4207578) | PDB MMDB KEGG UniProtKB/SwissProt B.MOAD antibodypedia GoogleScholar AffyNet | Purchase PC cid PC sid UniChem | Article PubMed | n/a | n/a | >5.00E+6 | n/a | n/a | n/a | n/a | n/a | n/a |
Jagiellonian University Curated by ChEMBL | Assay Description Inhibition of human USP2a (258 to 605 residues) expressed in Escherichia coli BL21(DE3) using Ub-AMC as substrate incubated for 30 mins measured at 3... | Eur J Med Chem 150: 261-267 (2018) Article DOI: 10.1016/j.ejmech.2018.03.009 BindingDB Entry DOI: 10.7270/Q28G8P98 | |||||||||||
| More data for this Ligand-Target Pair | |||||||||||||
| E3 ubiquitin-protein ligase Mdm2 [1-118] (Homo sapiens (Human)) | BDBM207985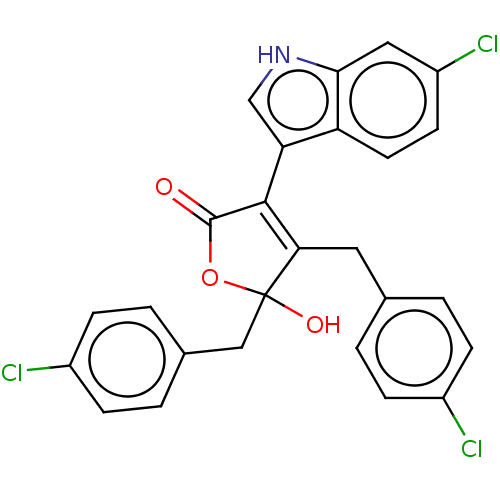 (4,5-di(4-chlorobenzyl)-3-(6-chloro-1H-indol-3-yl)-...) | PDB UniProtKB/SwissProt GoogleScholar AffyNet | PC cid PC sid UniChem | Article PubMed | n/a | n/a | n/a | 8.60E+4 | n/a | n/a | n/a | 7.4 | 15 |
Jagiellonian University | Assay Description Uniform 15N isotope labeling was achieved by expression of the protein in the M9 minimal media containing 15NH4Cl as the sole nitrogen source. The fi... | ACS Chem Biol 11: 3310-3318 (2016) Article DOI: 10.1021/acschembio.6b00596 BindingDB Entry DOI: 10.7270/Q2K35SH2 | |||||||||||
| More data for this Ligand-Target Pair | |||||||||||||
| E3 ubiquitin-protein ligase Mdm2 [1-118] (Homo sapiens (Human)) | BDBM207986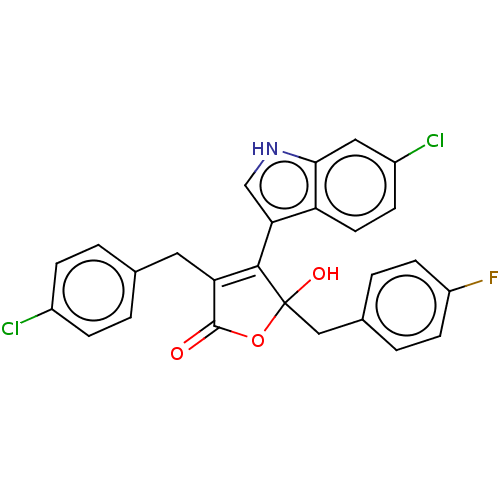 (3-(4-Chlorobenzyl)-4-(6-chloro-1H-indol-3-yl)-5-(4...) | PDB UniProtKB/SwissProt GoogleScholar AffyNet | PC cid PC sid UniChem | Article PubMed | n/a | n/a | n/a | 1.80E+4 | n/a | n/a | n/a | 7.4 | 15 |
Jagiellonian University | Assay Description Uniform 15N isotope labeling was achieved by expression of the protein in the M9 minimal media containing 15NH4Cl as the sole nitrogen source. The fi... | ACS Chem Biol 11: 3310-3318 (2016) Article DOI: 10.1021/acschembio.6b00596 BindingDB Entry DOI: 10.7270/Q2K35SH2 | |||||||||||
| More data for this Ligand-Target Pair | |||||||||||||
| E3 ubiquitin-protein ligase Mdm2 [1-118] (Homo sapiens (Human)) | BDBM207988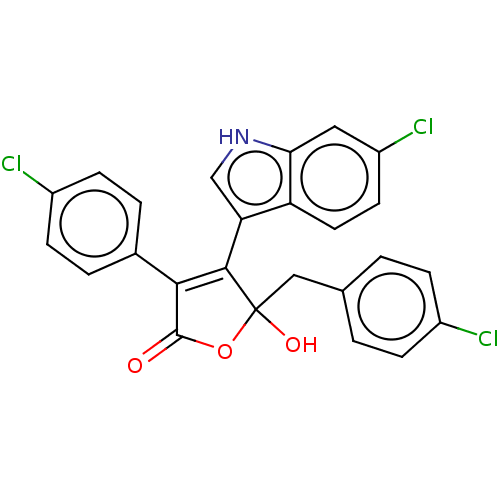 (5-(4-Chlorobenzyl)-4-(6-chloro-1H-indol-3-yl)-3-(4...) | PDB UniProtKB/SwissProt GoogleScholar AffyNet | PC cid PC sid UniChem | Article PubMed | n/a | n/a | n/a | 6.00E+4 | n/a | n/a | n/a | 7.4 | 15 |
Jagiellonian University | Assay Description Uniform 15N isotope labeling was achieved by expression of the protein in the M9 minimal media containing 15NH4Cl as the sole nitrogen source. The fi... | ACS Chem Biol 11: 3310-3318 (2016) Article DOI: 10.1021/acschembio.6b00596 BindingDB Entry DOI: 10.7270/Q2K35SH2 | |||||||||||
| More data for this Ligand-Target Pair | |||||||||||||
| E3 ubiquitin-protein ligase Mdm2 [1-118] (Homo sapiens (Human)) | BDBM207989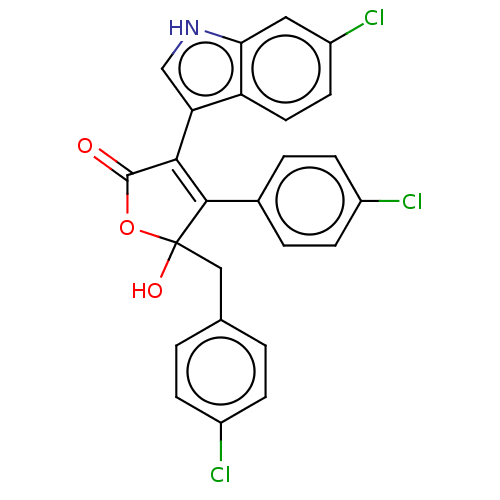 (5-(4-chlorobenzyl)-3-(6-chloro-1H-indol-3-yl)-4-(4...) | PDB UniProtKB/SwissProt GoogleScholar AffyNet | PC cid PC sid UniChem | Article PubMed | n/a | n/a | n/a | 6.80E+4 | n/a | n/a | n/a | 7.4 | 15 |
Jagiellonian University | Assay Description Uniform 15N isotope labeling was achieved by expression of the protein in the M9 minimal media containing 15NH4Cl as the sole nitrogen source. The fi... | ACS Chem Biol 11: 3310-3318 (2016) Article DOI: 10.1021/acschembio.6b00596 BindingDB Entry DOI: 10.7270/Q2K35SH2 | |||||||||||
| More data for this Ligand-Target Pair | |||||||||||||
| E3 ubiquitin-protein ligase Mdm2 [1-118,T47W] (Homo sapiens (Human)) | BDBM207976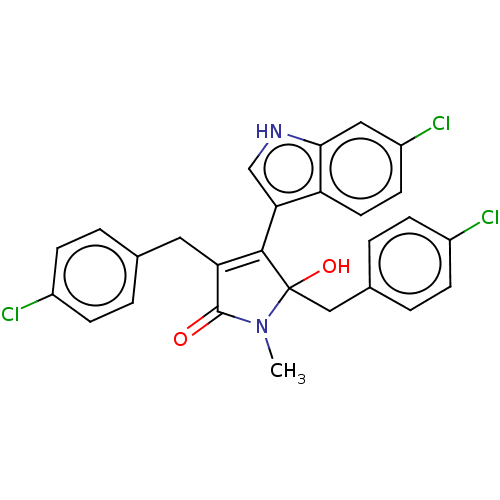 (N-Methyl-3,5-di(4-chlorobenzyl)-4-(6-chloro-1H-ind...) | PDB UniProtKB/SwissProt GoogleScholar AffyNet | PC cid PC sid PDB UniChem | PDB Article PubMed | n/a | n/a | n/a | 500 | n/a | n/a | n/a | 7.4 | 25 |
Jagiellonian University | Assay Description MST (NanoTemper Technologies GmbH) was used to determine the binding affinities between Mdm2 (residues 1−118 T47W; 500 nM) and inhibitors. T47W... | ACS Chem Biol 11: 3310-3318 (2016) Article DOI: 10.1021/acschembio.6b00596 BindingDB Entry DOI: 10.7270/Q2K35SH2 | |||||||||||
| More data for this Ligand-Target Pair |  3D Structure (crystal) | ||||||||||||
| E3 ubiquitin-protein ligase Mdm2 [1-118,T47W] (Homo sapiens (Human)) | BDBM207978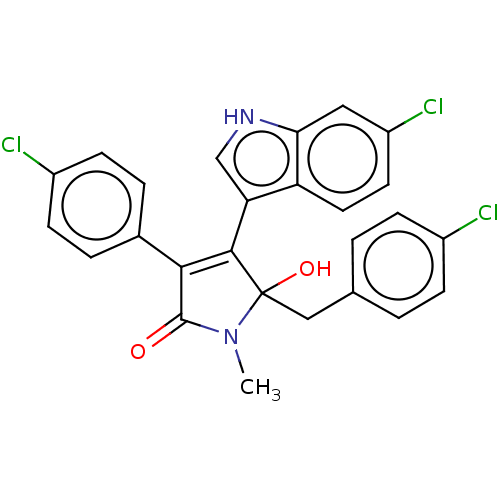 (N-Methyl-5-(4-chlorobenzyl)-4-(6-chloro-1H-indol-3...) | PDB UniProtKB/SwissProt GoogleScholar AffyNet | PC cid PC sid UniChem | Article PubMed | n/a | n/a | n/a | 1.87E+4 | n/a | n/a | n/a | 7.4 | 25 |
Jagiellonian University | Assay Description MST (NanoTemper Technologies GmbH) was used to determine the binding affinities between Mdm2 (residues 1−118 T47W; 500 nM) and inhibitors. T47W... | ACS Chem Biol 11: 3310-3318 (2016) Article DOI: 10.1021/acschembio.6b00596 BindingDB Entry DOI: 10.7270/Q2K35SH2 | |||||||||||
| More data for this Ligand-Target Pair | |||||||||||||
| E3 ubiquitin-protein ligase Mdm2 [1-118,T47W] (Homo sapiens (Human)) | BDBM207979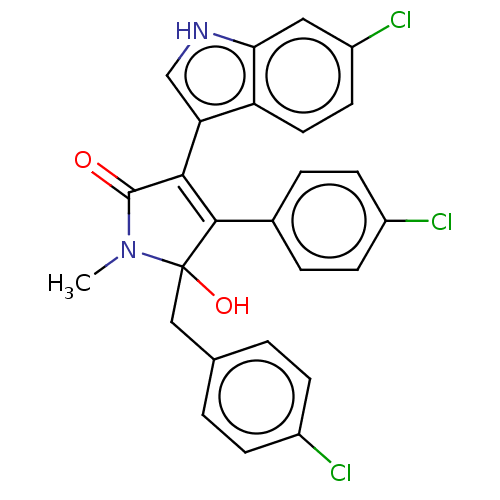 (N-methyl-5-(4-chlorobenzyl)-3-(6-chloro-1H-indol-3...) | PDB UniProtKB/SwissProt GoogleScholar AffyNet | PC cid PC sid UniChem | Article PubMed | n/a | n/a | n/a | 2.77E+4 | n/a | n/a | n/a | 7.4 | 25 |
Jagiellonian University | Assay Description MST (NanoTemper Technologies GmbH) was used to determine the binding affinities between Mdm2 (residues 1−118 T47W; 500 nM) and inhibitors. T47W... | ACS Chem Biol 11: 3310-3318 (2016) Article DOI: 10.1021/acschembio.6b00596 BindingDB Entry DOI: 10.7270/Q2K35SH2 | |||||||||||
| More data for this Ligand-Target Pair | |||||||||||||
| E3 ubiquitin-protein ligase Mdm2 [1-118,T47W] (Homo sapiens (Human)) | BDBM207980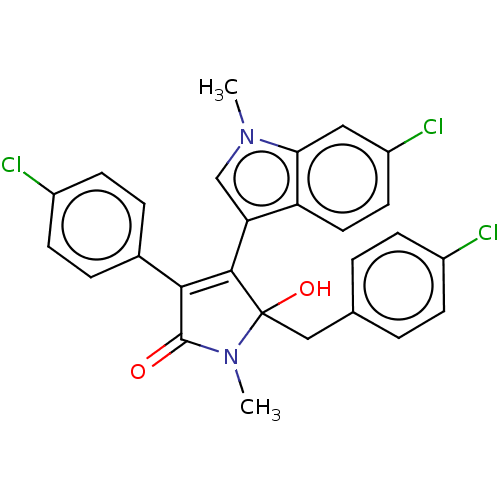 (N-Methyl-5-(4-chlorobenzyl)-4-(6-chloro-1-methyl-1...) | PDB UniProtKB/SwissProt GoogleScholar AffyNet | PC cid PC sid UniChem | Article PubMed | n/a | n/a | n/a | 4.05E+4 | n/a | n/a | n/a | 7.4 | 25 |
Jagiellonian University | Assay Description MST (NanoTemper Technologies GmbH) was used to determine the binding affinities between Mdm2 (residues 1−118 T47W; 500 nM) and inhibitors. T47W... | ACS Chem Biol 11: 3310-3318 (2016) Article DOI: 10.1021/acschembio.6b00596 BindingDB Entry DOI: 10.7270/Q2K35SH2 | |||||||||||
| More data for this Ligand-Target Pair | |||||||||||||
| E3 ubiquitin-protein ligase Mdm2 [1-118,T47W] (Homo sapiens (Human)) | BDBM207981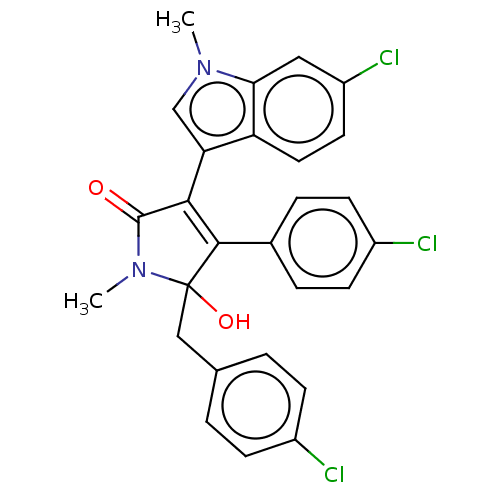 (N-methyl-5-(4-chlorobenzyl)-3-(6-chloro-1-methyl-1...) | PDB UniProtKB/SwissProt GoogleScholar AffyNet | PC cid PC sid UniChem | Article PubMed | n/a | n/a | n/a | 3.04E+4 | n/a | n/a | n/a | 7.4 | 25 |
Jagiellonian University | Assay Description MST (NanoTemper Technologies GmbH) was used to determine the binding affinities between Mdm2 (residues 1−118 T47W; 500 nM) and inhibitors. T47W... | ACS Chem Biol 11: 3310-3318 (2016) Article DOI: 10.1021/acschembio.6b00596 BindingDB Entry DOI: 10.7270/Q2K35SH2 | |||||||||||
| More data for this Ligand-Target Pair | |||||||||||||
| E3 ubiquitin-protein ligase Mdm2 [1-118,T47W] (Homo sapiens (Human)) | BDBM207982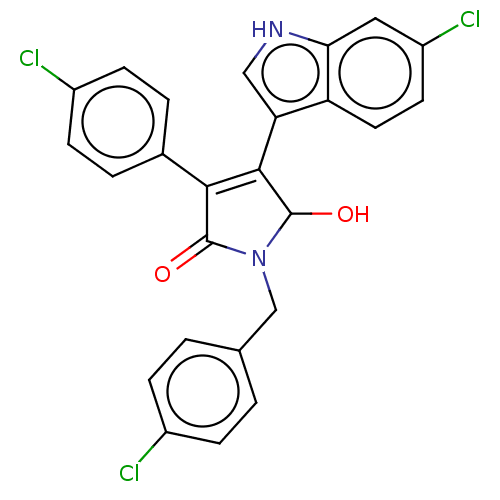 (N-(4-chlorobenzyl)-4-(6-chloro-1H-indol-3-yl)- 3-(...) | PDB UniProtKB/SwissProt GoogleScholar AffyNet | PC cid PC sid UniChem | Article PubMed | n/a | n/a | n/a | 2.44E+4 | n/a | n/a | n/a | 7.4 | 25 |
Jagiellonian University | Assay Description MST (NanoTemper Technologies GmbH) was used to determine the binding affinities between Mdm2 (residues 1−118 T47W; 500 nM) and inhibitors. T47W... | ACS Chem Biol 11: 3310-3318 (2016) Article DOI: 10.1021/acschembio.6b00596 BindingDB Entry DOI: 10.7270/Q2K35SH2 | |||||||||||
| More data for this Ligand-Target Pair | |||||||||||||
| E3 ubiquitin-protein ligase Mdm2 [1-118,T47W] (Homo sapiens (Human)) | BDBM207983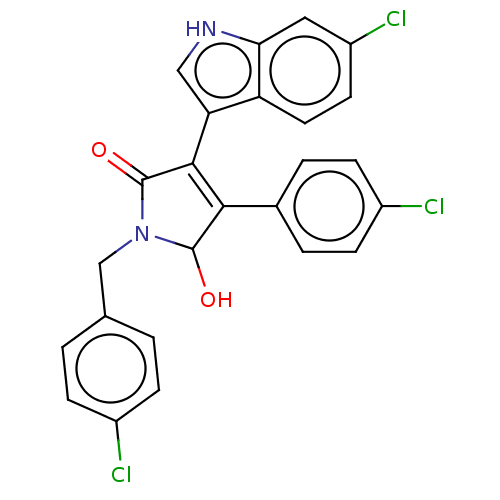 (N-(4-Chlorobenzyl)-3-(6-chloro-1H-indol-3-yl)-4-(4...) | PDB UniProtKB/SwissProt GoogleScholar AffyNet | PC cid PC sid UniChem | Article PubMed | n/a | n/a | n/a | 2.10E+3 | n/a | n/a | n/a | 7.4 | 25 |
Jagiellonian University | Assay Description MST (NanoTemper Technologies GmbH) was used to determine the binding affinities between Mdm2 (residues 1−118 T47W; 500 nM) and inhibitors. T47W... | ACS Chem Biol 11: 3310-3318 (2016) Article DOI: 10.1021/acschembio.6b00596 BindingDB Entry DOI: 10.7270/Q2K35SH2 | |||||||||||
| More data for this Ligand-Target Pair | |||||||||||||
| E3 ubiquitin-protein ligase Mdm2 [1-118,T47W] (Homo sapiens (Human)) | BDBM207984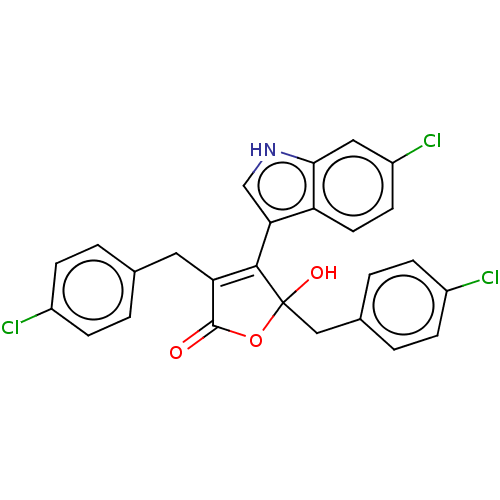 (3,5-Di(4-chlorobenzyl)-4-(6-chloro-1H-indol-3-yl)-...) | PDB UniProtKB/SwissProt GoogleScholar AffyNet | PC cid PC sid PDB UniChem | PDB Article PubMed | n/a | n/a | n/a | 3.10E+3 | n/a | n/a | n/a | 7.4 | 25 |
Jagiellonian University | Assay Description MST (NanoTemper Technologies GmbH) was used to determine the binding affinities between Mdm2 (residues 1−118 T47W; 500 nM) and inhibitors. T47W... | ACS Chem Biol 11: 3310-3318 (2016) Article DOI: 10.1021/acschembio.6b00596 BindingDB Entry DOI: 10.7270/Q2K35SH2 | |||||||||||
| More data for this Ligand-Target Pair |  3D Structure (crystal) | ||||||||||||
| E3 ubiquitin-protein ligase Mdm2 [1-118,T47W] (Homo sapiens (Human)) | BDBM207985 (4,5-di(4-chlorobenzyl)-3-(6-chloro-1H-indol-3-yl)-...) | PDB UniProtKB/SwissProt GoogleScholar AffyNet | PC cid PC sid UniChem | Article PubMed | n/a | n/a | n/a | 1.50E+3 | n/a | n/a | n/a | 7.4 | 25 |
Jagiellonian University | Assay Description MST (NanoTemper Technologies GmbH) was used to determine the binding affinities between Mdm2 (residues 1−118 T47W; 500 nM) and inhibitors. T47W... | ACS Chem Biol 11: 3310-3318 (2016) Article DOI: 10.1021/acschembio.6b00596 BindingDB Entry DOI: 10.7270/Q2K35SH2 | |||||||||||
| More data for this Ligand-Target Pair | |||||||||||||
| E3 ubiquitin-protein ligase Mdm2 [1-118,T47W] (Homo sapiens (Human)) | BDBM207987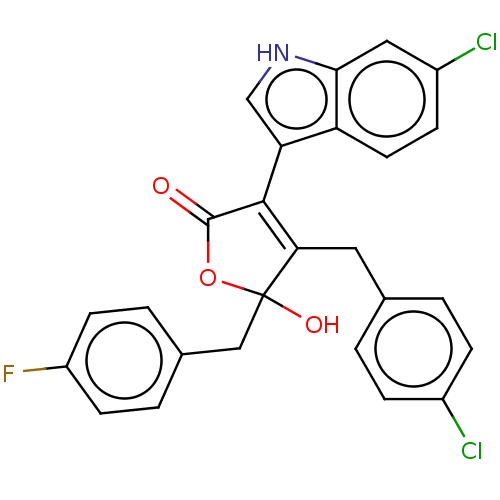 (4-(4-chlorobenzyl)-3-(6-chloro-1H-indol-3-yl)-5-(4...) | PDB UniProtKB/SwissProt GoogleScholar AffyNet | PC cid PC sid UniChem | Article PubMed | n/a | n/a | n/a | 2.60E+4 | n/a | n/a | n/a | 7.4 | 25 |
Jagiellonian University | Assay Description MST (NanoTemper Technologies GmbH) was used to determine the binding affinities between Mdm2 (residues 1−118 T47W; 500 nM) and inhibitors. T47W... | ACS Chem Biol 11: 3310-3318 (2016) Article DOI: 10.1021/acschembio.6b00596 BindingDB Entry DOI: 10.7270/Q2K35SH2 | |||||||||||
| More data for this Ligand-Target Pair | |||||||||||||
| E3 ubiquitin-protein ligase Mdm2 [1-118,T47W] (Homo sapiens (Human)) | BDBM207989 (5-(4-chlorobenzyl)-3-(6-chloro-1H-indol-3-yl)-4-(4...) | PDB UniProtKB/SwissProt GoogleScholar AffyNet | PC cid PC sid UniChem | Article PubMed | n/a | n/a | n/a | 7.00E+3 | n/a | n/a | n/a | 7.4 | 25 |
Jagiellonian University | Assay Description MST (NanoTemper Technologies GmbH) was used to determine the binding affinities between Mdm2 (residues 1−118 T47W; 500 nM) and inhibitors. T47W... | ACS Chem Biol 11: 3310-3318 (2016) Article DOI: 10.1021/acschembio.6b00596 BindingDB Entry DOI: 10.7270/Q2K35SH2 | |||||||||||
| More data for this Ligand-Target Pair | |||||||||||||
| E3 ubiquitin-protein ligase Mdm2 [1-118,T47W] (Homo sapiens (Human)) | BDBM207986 (3-(4-Chlorobenzyl)-4-(6-chloro-1H-indol-3-yl)-5-(4...) | PDB UniProtKB/SwissProt GoogleScholar AffyNet | PC cid PC sid UniChem | Article PubMed | n/a | n/a | n/a | 2.87E+4 | n/a | n/a | n/a | 7.4 | 25 |
Jagiellonian University | Assay Description MST (NanoTemper Technologies GmbH) was used to determine the binding affinities between Mdm2 (residues 1−118 T47W; 500 nM) and inhibitors. T47W... | ACS Chem Biol 11: 3310-3318 (2016) Article DOI: 10.1021/acschembio.6b00596 BindingDB Entry DOI: 10.7270/Q2K35SH2 | |||||||||||
| More data for this Ligand-Target Pair | |||||||||||||
| E3 ubiquitin-protein ligase Mdm2 [1-118,T47W] (Homo sapiens (Human)) | BDBM207988 (5-(4-Chlorobenzyl)-4-(6-chloro-1H-indol-3-yl)-3-(4...) | PDB UniProtKB/SwissProt GoogleScholar AffyNet | PC cid PC sid UniChem | Article PubMed | n/a | n/a | n/a | 1.30E+3 | n/a | n/a | n/a | 7.4 | 25 |
Jagiellonian University | Assay Description MST (NanoTemper Technologies GmbH) was used to determine the binding affinities between Mdm2 (residues 1−118 T47W; 500 nM) and inhibitors. T47W... | ACS Chem Biol 11: 3310-3318 (2016) Article DOI: 10.1021/acschembio.6b00596 BindingDB Entry DOI: 10.7270/Q2K35SH2 | |||||||||||
| More data for this Ligand-Target Pair | |||||||||||||
| Ubiquitin carboxyl-terminal hydrolase 2 (Homo sapiens (Human)) | BDBM50458416 (CHEMBL1230155) | PDB MMDB KEGG UniProtKB/SwissProt B.MOAD antibodypedia GoogleScholar AffyNet | Purchase MCE PC cid PC sid PDB UniChem | Article PubMed | n/a | n/a | n/a | 9.00E+5 | n/a | n/a | n/a | n/a | n/a |
Jagiellonian University Curated by ChEMBL | Assay Description Binding affinity to 15N-Gly/Cys/Ser/Trp-labeled human USP2a expressed in Escherichia coli BL21(DE3) assessed as change in chemical shift by NMR metho... | Eur J Med Chem 150: 261-267 (2018) Article DOI: 10.1016/j.ejmech.2018.03.009 BindingDB Entry DOI: 10.7270/Q28G8P98 | |||||||||||
| More data for this Ligand-Target Pair | |||||||||||||
| E3 ubiquitin-protein ligase Mdm2 [1-118] (Homo sapiens (Human)) | BDBM207984 (3,5-Di(4-chlorobenzyl)-4-(6-chloro-1H-indol-3-yl)-...) | PDB UniProtKB/SwissProt GoogleScholar AffyNet | PC cid PC sid PDB UniChem | PDB Article PubMed | n/a | n/a | n/a | <1.00E+3 | n/a | n/a | n/a | 7.4 | 15 |
Jagiellonian University | Assay Description Uniform 15N isotope labeling was achieved by expression of the protein in the M9 minimal media containing 15NH4Cl as the sole nitrogen source. The fi... | ACS Chem Biol 11: 3310-3318 (2016) Article DOI: 10.1021/acschembio.6b00596 BindingDB Entry DOI: 10.7270/Q2K35SH2 | |||||||||||
| More data for this Ligand-Target Pair |  3D Structure (crystal) | ||||||||||||
| E3 ubiquitin-protein ligase Mdm2 [1-118] (Homo sapiens (Human)) | BDBM207983 (N-(4-Chlorobenzyl)-3-(6-chloro-1H-indol-3-yl)-4-(4...) | PDB UniProtKB/SwissProt GoogleScholar AffyNet | PC cid PC sid UniChem | Article PubMed | n/a | n/a | n/a | 4.00E+4 | n/a | n/a | n/a | 7.4 | 15 |
Jagiellonian University | Assay Description Uniform 15N isotope labeling was achieved by expression of the protein in the M9 minimal media containing 15NH4Cl as the sole nitrogen source. The fi... | ACS Chem Biol 11: 3310-3318 (2016) Article DOI: 10.1021/acschembio.6b00596 BindingDB Entry DOI: 10.7270/Q2K35SH2 | |||||||||||
| More data for this Ligand-Target Pair | |||||||||||||
| E3 ubiquitin-protein ligase Mdm2 [1-118] (Homo sapiens (Human)) | BDBM207982 (N-(4-chlorobenzyl)-4-(6-chloro-1H-indol-3-yl)- 3-(...) | PDB UniProtKB/SwissProt GoogleScholar AffyNet | PC cid PC sid UniChem | Article PubMed | n/a | n/a | n/a | 1.64E+5 | n/a | n/a | n/a | 7.4 | 15 |
Jagiellonian University | Assay Description Uniform 15N isotope labeling was achieved by expression of the protein in the M9 minimal media containing 15NH4Cl as the sole nitrogen source. The fi... | ACS Chem Biol 11: 3310-3318 (2016) Article DOI: 10.1021/acschembio.6b00596 BindingDB Entry DOI: 10.7270/Q2K35SH2 | |||||||||||
| More data for this Ligand-Target Pair | |||||||||||||
| E3 ubiquitin-protein ligase Mdm2 [1-118] (Homo sapiens (Human)) | BDBM207981 (N-methyl-5-(4-chlorobenzyl)-3-(6-chloro-1-methyl-1...) | PDB UniProtKB/SwissProt GoogleScholar AffyNet | PC cid PC sid UniChem | Article PubMed | n/a | n/a | n/a | 2.46E+5 | n/a | n/a | n/a | 7.4 | 15 |
Jagiellonian University | Assay Description Uniform 15N isotope labeling was achieved by expression of the protein in the M9 minimal media containing 15NH4Cl as the sole nitrogen source. The fi... | ACS Chem Biol 11: 3310-3318 (2016) Article DOI: 10.1021/acschembio.6b00596 BindingDB Entry DOI: 10.7270/Q2K35SH2 | |||||||||||
| More data for this Ligand-Target Pair | |||||||||||||
| E3 ubiquitin-protein ligase Mdm2 [1-118] (Homo sapiens (Human)) | BDBM207980 (N-Methyl-5-(4-chlorobenzyl)-4-(6-chloro-1-methyl-1...) | PDB UniProtKB/SwissProt GoogleScholar AffyNet | PC cid PC sid UniChem | Article PubMed | n/a | n/a | n/a | 3.65E+5 | n/a | n/a | n/a | 7.4 | 15 |
Jagiellonian University | Assay Description Uniform 15N isotope labeling was achieved by expression of the protein in the M9 minimal media containing 15NH4Cl as the sole nitrogen source. The fi... | ACS Chem Biol 11: 3310-3318 (2016) Article DOI: 10.1021/acschembio.6b00596 BindingDB Entry DOI: 10.7270/Q2K35SH2 | |||||||||||
| More data for this Ligand-Target Pair | |||||||||||||
| E3 ubiquitin-protein ligase Mdm2 [1-118] (Homo sapiens (Human)) | BDBM207979 (N-methyl-5-(4-chlorobenzyl)-3-(6-chloro-1H-indol-3...) | PDB UniProtKB/SwissProt GoogleScholar AffyNet | PC cid PC sid UniChem | Article PubMed | n/a | n/a | n/a | 1.92E+5 | n/a | n/a | n/a | 7.4 | 15 |
Jagiellonian University | Assay Description Uniform 15N isotope labeling was achieved by expression of the protein in the M9 minimal media containing 15NH4Cl as the sole nitrogen source. The fi... | ACS Chem Biol 11: 3310-3318 (2016) Article DOI: 10.1021/acschembio.6b00596 BindingDB Entry DOI: 10.7270/Q2K35SH2 | |||||||||||
| More data for this Ligand-Target Pair | |||||||||||||
| E3 ubiquitin-protein ligase Mdm2 [1-118] (Homo sapiens (Human)) | BDBM207978 (N-Methyl-5-(4-chlorobenzyl)-4-(6-chloro-1H-indol-3...) | PDB UniProtKB/SwissProt GoogleScholar AffyNet | PC cid PC sid UniChem | Article PubMed | n/a | n/a | n/a | 5.10E+4 | n/a | n/a | n/a | 7.4 | 15 |
Jagiellonian University | Assay Description Uniform 15N isotope labeling was achieved by expression of the protein in the M9 minimal media containing 15NH4Cl as the sole nitrogen source. The fi... | ACS Chem Biol 11: 3310-3318 (2016) Article DOI: 10.1021/acschembio.6b00596 BindingDB Entry DOI: 10.7270/Q2K35SH2 | |||||||||||
| More data for this Ligand-Target Pair | |||||||||||||
| E3 ubiquitin-protein ligase Mdm2 [1-118] (Homo sapiens (Human)) | BDBM207976 (N-Methyl-3,5-di(4-chlorobenzyl)-4-(6-chloro-1H-ind...) | PDB UniProtKB/SwissProt GoogleScholar AffyNet | PC cid PC sid PDB UniChem | PDB Article PubMed | n/a | n/a | n/a | <1.00E+3 | n/a | n/a | n/a | 7.4 | 15 |
Jagiellonian University | Assay Description Uniform 15N isotope labeling was achieved by expression of the protein in the M9 minimal media containing 15NH4Cl as the sole nitrogen source. The fi... | ACS Chem Biol 11: 3310-3318 (2016) Article DOI: 10.1021/acschembio.6b00596 BindingDB Entry DOI: 10.7270/Q2K35SH2 | |||||||||||
| More data for this Ligand-Target Pair |  3D Structure (crystal) | ||||||||||||
| E3 ubiquitin-protein ligase Mdm2 [1-118] (Homo sapiens (Human)) | BDBM207977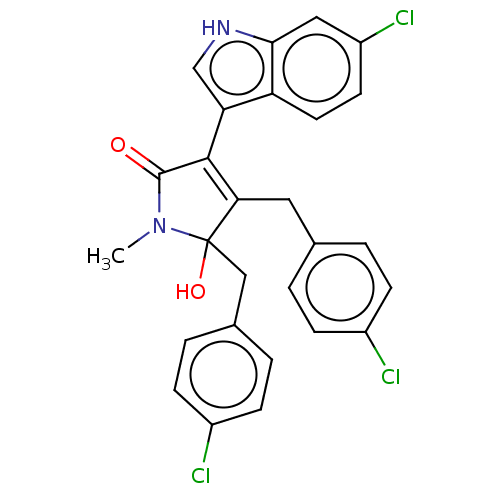 (N-methyl-4,5-di(4-chlorobenzyl)-3-(6-chloro-1H-ind...) | PDB UniProtKB/SwissProt GoogleScholar AffyNet | PC cid PC sid UniChem | Article PubMed | n/a | n/a | n/a | 5.60E+4 | n/a | n/a | n/a | 7.4 | 15 |
Jagiellonian University | Assay Description Uniform 15N isotope labeling was achieved by expression of the protein in the M9 minimal media containing 15NH4Cl as the sole nitrogen source. The fi... | ACS Chem Biol 11: 3310-3318 (2016) Article DOI: 10.1021/acschembio.6b00596 BindingDB Entry DOI: 10.7270/Q2K35SH2 | |||||||||||
| More data for this Ligand-Target Pair | |||||||||||||