| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Poly [ADP-ribose] polymerase 2 |
|---|
| Ligand | BDBM50593627 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_2268421 |
|---|
| IC50 | 200±n/a nM |
|---|
| Citation |  Buchstaller, HP; Anlauf, U; Dorsch, D; K�gler, S; Kuhn, D; Lehmann, M; Leuthner, B; Lodholz, S; Musil, D; Radtki, D; Rettig, C; Ritzert, C; Rohdich, F; Schneider, R; Wegener, A; Weigt, S; Wilkinson, K; Esdar, C Optimization of a Screening Hit toward M2912, an Oral Tankyrase Inhibitor with Antitumor Activity in Colorectal Cancer Models. J Med Chem64:10371-10392 (2021) [PubMed] Buchstaller, HP; Anlauf, U; Dorsch, D; K�gler, S; Kuhn, D; Lehmann, M; Leuthner, B; Lodholz, S; Musil, D; Radtki, D; Rettig, C; Ritzert, C; Rohdich, F; Schneider, R; Wegener, A; Weigt, S; Wilkinson, K; Esdar, C Optimization of a Screening Hit toward M2912, an Oral Tankyrase Inhibitor with Antitumor Activity in Colorectal Cancer Models. J Med Chem64:10371-10392 (2021) [PubMed] |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Poly [ADP-ribose] polymerase 2 |
|---|
| Name: | Poly [ADP-ribose] polymerase 2 |
|---|
| Synonyms: | (ARTD2 or PARP2) | ADPRT2 | ADPRTL2 | PARP2 | PARP2_HUMAN | Poly [ADP-ribose] polymerase 2 (PARP-2) | Poly [ADP-ribose] polymerase 2 (PARP2) |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 66225.70 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | Q9UGN5 |
|---|
| Residue: | 583 |
|---|
| Sequence: | MAARRRRSTGGGRARALNESKRVNNGNTAPEDSSPAKKTRRCQRQESKKMPVAGGKANKD
RTEDKQDGMPGRSWASKRVSESVKALLLKGKAPVDPECTAKVGKAHVYCEGNDVYDVMLN
QTNLQFNNNKYYLIQLLEDDAQRNFSVWMRWGRVGKMGQHSLVACSGNLNKAKEIFQKKF
LDKTKNNWEDREKFEKVPGKYDMLQMDYATNTQDEEETKKEESLKSPLKPESQLDLRVQE
LIKLICNVQAMEEMMMEMKYNTKKAPLGKLTVAQIKAGYQSLKKIEDCIRAGQHGRALME
ACNEFYTRIPHDFGLRTPPLIRTQKELSEKIQLLEALGDIEIAIKLVKTELQSPEHPLDQ
HYRNLHCALRPLDHESYEFKVISQYLQSTHAPTHSDYTMTLLDLFEVEKDGEKEAFREDL
HNRMLLWHGSRMSNWVGILSHGLRIAPPEAPITGYMFGKGIYFADMSSKSANYCFASRLK
NTGLLLLSEVALGQCNELLEANPKAEGLLQGKHSTKGLGKMAPSSAHFVTLNGSTVPLGP
ASDTGILNPDGYTLNYNEYIVYNPNQVRMRYLLKVQFNFLQLW
|
|
|
|---|
| BDBM50593627 |
|---|
| n/a |
|---|
| Name | BDBM50593627 |
|---|
| Synonyms: | CHEMBL5199558 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C17H18N2O2 |
|---|
| Mol. Mass. | 282.337 |
|---|
| SMILES | Cc1ccc2n1cc([nH]c2=O)-c1ccc(cc1)C(C)(C)O |
|---|
| Structure | 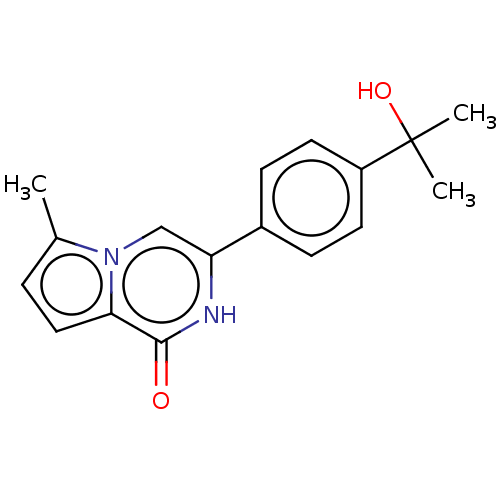
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Buchstaller, HP; Anlauf, U; Dorsch, D; K�gler, S; Kuhn, D; Lehmann, M; Leuthner, B; Lodholz, S; Musil, D; Radtki, D; Rettig, C; Ritzert, C; Rohdich, F; Schneider, R; Wegener, A; Weigt, S; Wilkinson, K; Esdar, C Optimization of a Screening Hit toward M2912, an Oral Tankyrase Inhibitor with Antitumor Activity in Colorectal Cancer Models. J Med Chem64:10371-10392 (2021) [PubMed]
Buchstaller, HP; Anlauf, U; Dorsch, D; K�gler, S; Kuhn, D; Lehmann, M; Leuthner, B; Lodholz, S; Musil, D; Radtki, D; Rettig, C; Ritzert, C; Rohdich, F; Schneider, R; Wegener, A; Weigt, S; Wilkinson, K; Esdar, C Optimization of a Screening Hit toward M2912, an Oral Tankyrase Inhibitor with Antitumor Activity in Colorectal Cancer Models. J Med Chem64:10371-10392 (2021) [PubMed]