| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Beta-1 adrenergic receptor |
|---|
| Ligand | BDBM50338815 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_727080 (CHEMBL1687004) |
|---|
| IC50 | 2930±n/a nM |
|---|
| Citation |  Morriello, GJ; Wendt, HR; Bansal, A; Di Salvo, J; Feighner, S; He, J; Hurley, AL; Hreniuk, DL; Salituro, GM; Reddy, MV; Galloway, SM; McGettigan, KK; Laws, G; McKnight, C; Doss, GA; Tsou, NN; Black, RM; Morris, J; Ball, RG; Sanfiz, AT; Streckfuss, E; Struthers, M; Edmondson, SD Design of a novel pyrrolidine scaffold utilized in the discovery of potent and selective humanß3 adrenergic receptor agonists. Bioorg Med Chem Lett21:1865-70 (2011) [PubMed] Article Morriello, GJ; Wendt, HR; Bansal, A; Di Salvo, J; Feighner, S; He, J; Hurley, AL; Hreniuk, DL; Salituro, GM; Reddy, MV; Galloway, SM; McGettigan, KK; Laws, G; McKnight, C; Doss, GA; Tsou, NN; Black, RM; Morris, J; Ball, RG; Sanfiz, AT; Streckfuss, E; Struthers, M; Edmondson, SD Design of a novel pyrrolidine scaffold utilized in the discovery of potent and selective humanß3 adrenergic receptor agonists. Bioorg Med Chem Lett21:1865-70 (2011) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Beta-1 adrenergic receptor |
|---|
| Name: | Beta-1 adrenergic receptor |
|---|
| Synonyms: | ADRB1 | ADRB1R | ADRB1_HUMAN | B1AR | Beta-1 adrenoceptor | Beta-1 adrenoreceptor | adrenergic Beta1 |
|---|
| Type: | G Protein-Coupled Receptor (GPCR) |
|---|
| Mol. Mass.: | 51338.40 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | P08588 |
|---|
| Residue: | 477 |
|---|
| Sequence: | MGAGVLVLGASEPGNLSSAAPLPDGAATAARLLVPASPPASLLPPASESPEPLSQQWTAG
MGLLMALIVLLIVAGNVLVIVAIAKTPRLQTLTNLFIMSLASADLVMGLLVVPFGATIVV
WGRWEYGSFFCELWTSVDVLCVTASIETLCVIALDRYLAITSPFRYQSLLTRARARGLVC
TVWAISALVSFLPILMHWWRAESDEARRCYNDPKCCDFVTNRAYAIASSVVSFYVPLCIM
AFVYLRVFREAQKQVKKIDSCERRFLGGPARPPSPSPSPVPAPAPPPGPPRPAAAAATAP
LANGRAGKRRPSRLVALREQKALKTLGIIMGVFTLCWLPFFLANVVKAFHRELVPDRLFV
FFNWLGYANSAFNPIIYCRSPDFRKAFQGLLCCARRAARRRHATHGDRPRASGCLARPGP
PPSPGAASDDDDDDVVGATPPARLLEPWAGCNGGAAADSDSSLDEPCRPGFASESKV
|
|
|
|---|
| BDBM50338815 |
|---|
| n/a |
|---|
| Name | BDBM50338815 |
|---|
| Synonyms: | CHEMBL1684577 | N-(4-(((5R)-5-((R)-hydroxy(phenyl)methyl)pyrrolidin-2-yl)methyl)phenyl)-4-(4-(4-(trifluoromethyl)phenyl)thiazol-2-yl)benzenesulfonamide |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C34H30F3N3O3S2 |
|---|
| Mol. Mass. | 649.746 |
|---|
| SMILES | O[C@@H]([C@H]1CCC(Cc2ccc(NS(=O)(=O)c3ccc(cc3)-c3nc(cs3)-c3ccc(cc3)C(F)(F)F)cc2)N1)c1ccccc1 |r| |
|---|
| Structure | 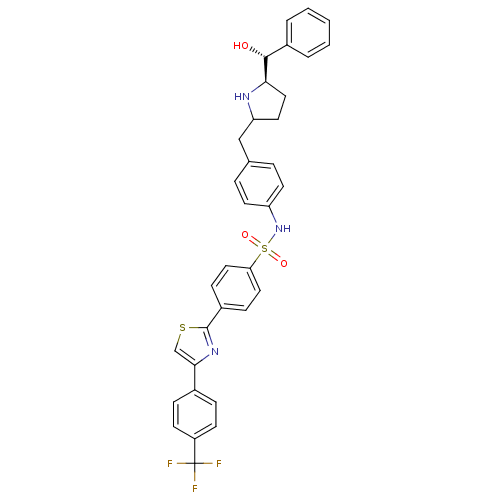
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Morriello, GJ; Wendt, HR; Bansal, A; Di Salvo, J; Feighner, S; He, J; Hurley, AL; Hreniuk, DL; Salituro, GM; Reddy, MV; Galloway, SM; McGettigan, KK; Laws, G; McKnight, C; Doss, GA; Tsou, NN; Black, RM; Morris, J; Ball, RG; Sanfiz, AT; Streckfuss, E; Struthers, M; Edmondson, SD Design of a novel pyrrolidine scaffold utilized in the discovery of potent and selective humanß3 adrenergic receptor agonists. Bioorg Med Chem Lett21:1865-70 (2011) [PubMed] Article
Morriello, GJ; Wendt, HR; Bansal, A; Di Salvo, J; Feighner, S; He, J; Hurley, AL; Hreniuk, DL; Salituro, GM; Reddy, MV; Galloway, SM; McGettigan, KK; Laws, G; McKnight, C; Doss, GA; Tsou, NN; Black, RM; Morris, J; Ball, RG; Sanfiz, AT; Streckfuss, E; Struthers, M; Edmondson, SD Design of a novel pyrrolidine scaffold utilized in the discovery of potent and selective humanß3 adrenergic receptor agonists. Bioorg Med Chem Lett21:1865-70 (2011) [PubMed] Article