| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Prothrombin |
|---|
| Ligand | BDBM18982 |
|---|
| Substrate/Competitor | BDBM13573 |
|---|
| Meas. Tech. | Enzyme Assay and Determination of the Inhibition Constants. |
|---|
| pH | 7±n/a |
|---|
| Temperature | 295.15±n/a K |
|---|
| Ki | 6500±n/a nM |
|---|
| Citation |  Lam, PY; Clark, CG; Li, R; Pinto, DJ; Orwat, MJ; Galemmo, RA; Fevig, JM; Teleha, CA; Alexander, RS; Smallwood, AM; Rossi, KA; Wright, MR; Bai, SA; He, K; Luettgen, JM; Wong, PC; Knabb, RM; Wexler, RR Structure-based design of novel guanidine/benzamidine mimics: potent and orally bioavailable factor Xa inhibitors as novel anticoagulants. J Med Chem46:4405-18 (2003) [PubMed] Article Lam, PY; Clark, CG; Li, R; Pinto, DJ; Orwat, MJ; Galemmo, RA; Fevig, JM; Teleha, CA; Alexander, RS; Smallwood, AM; Rossi, KA; Wright, MR; Bai, SA; He, K; Luettgen, JM; Wong, PC; Knabb, RM; Wexler, RR Structure-based design of novel guanidine/benzamidine mimics: potent and orally bioavailable factor Xa inhibitors as novel anticoagulants. J Med Chem46:4405-18 (2003) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Solution Info, Assay Method |
|---|
| |
| Prothrombin |
|---|
| Name: | Prothrombin |
|---|
| Synonyms: | Activation peptide fragment 1 | Activation peptide fragment 2 | Coagulation factor II | F2 | Prothrombin precursor | THRB_HUMAN | Thrombin heavy chain | Thrombin light chain |
|---|
| Type: | Protein |
|---|
| Mol. Mass.: | 70029.57 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | P00734 |
|---|
| Residue: | 622 |
|---|
| Sequence: | MAHVRGLQLPGCLALAALCSLVHSQHVFLAPQQARSLLQRVRRANTFLEEVRKGNLEREC
VEETCSYEEAFEALESSTATDVFWAKYTACETARTPRDKLAACLEGNCAEGLGTNYRGHV
NITRSGIECQLWRSRYPHKPEINSTTHPGADLQENFCRNPDSSTTGPWCYTTDPTVRRQE
CSIPVCGQDQVTVAMTPRSEGSSVNLSPPLEQCVPDRGQQYQGRLAVTTHGLPCLAWASA
QAKALSKHQDFNSAVQLVENFCRNPDGDEEGVWCYVAGKPGDFGYCDLNYCEEAVEEETG
DGLDEDSDRAIEGRTATSEYQTFFNPRTFGSGEADCGLRPLFEKKSLEDKTERELLESYI
DGRIVEGSDAEIGMSPWQVMLFRKSPQELLCGASLISDRWVLTAAHCLLYPPWDKNFTEN
DLLVRIGKHSRTRYERNIEKISMLEKIYIHPRYNWRENLDRDIALMKLKKPVAFSDYIHP
VCLPDRETAASLLQAGYKGRVTGWGNLKETWTANVGKGQPSVLQVVNLPIVERPVCKDST
RIRITDNMFCAGYKPDEGKRGDACEGDSGGPFVMKSPFNNRWYQMGIVSWGEGCDRDGKY
GFYTHVFRLKKWIQKVIDQFGE
|
|
|
|---|
| BDBM18982 |
|---|
| BDBM13573 |
|---|
| Name | BDBM18982 |
|---|
| Synonyms: | 1-(1-aminoisoquinolin-7-yl)-3-methyl-N-[4-(2-sulfamoylphenyl)phenyl]-1H-pyrazole-5-carboxamide | SQ-311 | pyrazole-based inhibitor, 24a |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C26H22N6O3S |
|---|
| Mol. Mass. | 498.556 |
|---|
| SMILES | Cc1cc(C(=O)Nc2ccc(cc2)-c2ccccc2S(N)(=O)=O)n(n1)-c1ccc2ccnc(N)c2c1 |
|---|
| Structure | 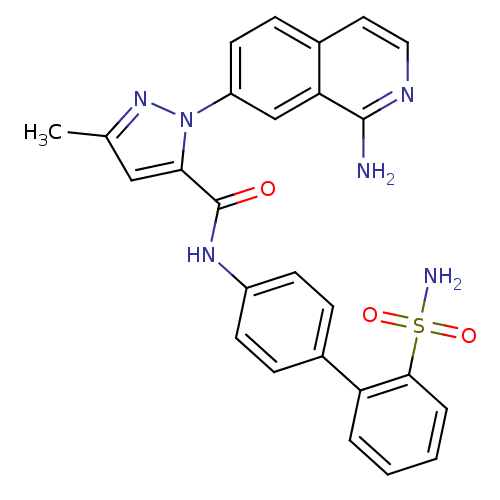
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Lam, PY; Clark, CG; Li, R; Pinto, DJ; Orwat, MJ; Galemmo, RA; Fevig, JM; Teleha, CA; Alexander, RS; Smallwood, AM; Rossi, KA; Wright, MR; Bai, SA; He, K; Luettgen, JM; Wong, PC; Knabb, RM; Wexler, RR Structure-based design of novel guanidine/benzamidine mimics: potent and orally bioavailable factor Xa inhibitors as novel anticoagulants. J Med Chem46:4405-18 (2003) [PubMed] Article
Lam, PY; Clark, CG; Li, R; Pinto, DJ; Orwat, MJ; Galemmo, RA; Fevig, JM; Teleha, CA; Alexander, RS; Smallwood, AM; Rossi, KA; Wright, MR; Bai, SA; He, K; Luettgen, JM; Wong, PC; Knabb, RM; Wexler, RR Structure-based design of novel guanidine/benzamidine mimics: potent and orally bioavailable factor Xa inhibitors as novel anticoagulants. J Med Chem46:4405-18 (2003) [PubMed] Article