| Reaction Details |
|---|
 | Report a problem with these data |
| Target | cAMP-dependent protein kinase catalytic subunit alpha |
|---|
| Ligand | BDBM27226 |
|---|
| Substrate/Competitor | BDBM27221 |
|---|
| Meas. Tech. | Fluorescence Polarization-Based Binding Displacement Kinase Assay |
|---|
| pH | 7.5±n/a |
|---|
| Temperature | 303.15±n/a K |
|---|
| Kd | 0.5±n/a nM |
|---|
| Citation |  Lavogina, D; Lust, M; Viil, I; König, N; Raidaru, G; Rogozina, J; Enkvist, E; Uri, A; Bossemeyer, D Structural analysis of ARC-type inhibitor (ARC-1034) binding to protein kinase A catalytic subunit and rational design of bisubstrate analogue inhibitors of basophilic protein kinases. J Med Chem52:308-21 (2009) [PubMed] Article Lavogina, D; Lust, M; Viil, I; König, N; Raidaru, G; Rogozina, J; Enkvist, E; Uri, A; Bossemeyer, D Structural analysis of ARC-type inhibitor (ARC-1034) binding to protein kinase A catalytic subunit and rational design of bisubstrate analogue inhibitors of basophilic protein kinases. J Med Chem52:308-21 (2009) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Inhibition_Run data, Solution Info, Assay Method |
|---|
| |
| cAMP-dependent protein kinase catalytic subunit alpha |
|---|
| Name: | cAMP-dependent protein kinase catalytic subunit alpha |
|---|
| Synonyms: | KAPCA_MOUSE | PKA C-alpha | Pkaca | Prkaca | cAMP-Dependent Protein Kinase (PKA) | cAMP-dependent protein kinase catalytic subunit alpha | cAMP-dependent protein kinase, alpha-catalytic subunit |
|---|
| Type: | Enzyme Catalytic Unit |
|---|
| Mol. Mass.: | 40579.18 |
|---|
| Organism: | Mus musculus (mouse) |
|---|
| Description: | n/a |
|---|
| Residue: | 351 |
|---|
| Sequence: | MGNAAAAKKGSEQESVKEFLAKAKEDFLKKWETPSQNTAQLDQFDRIKTLGTGSFGRVML
VKHKESGNHYAMKILDKQKVVKLKQIEHTLNEKRILQAVNFPFLVKLEFSFKDNSNLYMV
MEYVAGGEMFSHLRRIGRFSEPHARFYAAQIVLTFEYLHSLDLIYRDLKPENLLIDQQGY
IQVTDFGFAKRVKGRTWTLCGTPEYLAPEIILSKGYNKAVDWWALGVLIYEMAAGYPPFF
ADQPIQIYEKIVSGKVRFPSHFSSDLKDLLRNLLQVDLTKRFGNLKNGVNDIKNHKWFAT
TDWIAIYQRKVEAPFIPKFKGPGDTSNFDDYEEEEIRVSINEKCGKEFTEF
|
|
|
|---|
| BDBM27226 |
|---|
| BDBM27221 |
|---|
| Name | BDBM27226 |
|---|
| Synonyms: | (2R)-6-amino-2-(6-{[(2S,3S,4R,5R)-5-(6-amino-9H-purin-9-yl)-3,4-dihydroxyoxolan-2-yl]formamido}hexanamido)-N-(5-{[(1R)-4-carbamimidamido-1-{[(1R)-4-carbamimidamido-1-{[(1R)-4-carbamimidamido-1-{[(1R)-4-carbamimidamido-1-carbamoylbutyl]carbamoyl}butyl]carbamoyl}butyl]carbamoyl}butyl]carbamoyl}pentyl)hexanamide | ARC-1027, 21 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C52H94N26O11 |
|---|
| Mol. Mass. | 1259.4704 |
|---|
| SMILES | [#7]-[#6]-[#6]-[#6]-[#6]-[#6@@H](-[#7]-[#6](=O)-[#6]-[#6]-[#6]-[#6]-[#6]-[#7]-[#6](=O)-[#6@H]-1-[#8]-[#6@H](-[#6@H](-[#8])-[#6@@H]-1-[#8])-n1cnc2c(-[#7])ncnc12)-[#6](=O)-[#7]-[#6]-[#6]-[#6]-[#6]-[#6]-[#6](=O)-[#7]-[#6@H](-[#6]-[#6]-[#6]\[#7]=[#6](/[#7])-[#7])-[#6](=O)-[#7]-[#6@H](-[#6]-[#6]-[#6]\[#7]=[#6](\[#7])-[#7])-[#6](=O)-[#7]-[#6@H](-[#6]-[#6]-[#6]\[#7]=[#6](/[#7])-[#7])-[#6](=O)-[#7]-[#6@H](-[#6]-[#6]-[#6]\[#7]=[#6](\[#7])-[#7])-[#6](-[#7])=O |r| |
|---|
| Structure | 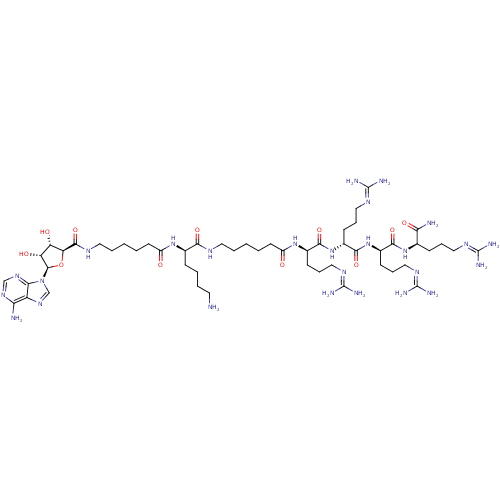
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Lavogina, D; Lust, M; Viil, I; König, N; Raidaru, G; Rogozina, J; Enkvist, E; Uri, A; Bossemeyer, D Structural analysis of ARC-type inhibitor (ARC-1034) binding to protein kinase A catalytic subunit and rational design of bisubstrate analogue inhibitors of basophilic protein kinases. J Med Chem52:308-21 (2009) [PubMed] Article
Lavogina, D; Lust, M; Viil, I; König, N; Raidaru, G; Rogozina, J; Enkvist, E; Uri, A; Bossemeyer, D Structural analysis of ARC-type inhibitor (ARC-1034) binding to protein kinase A catalytic subunit and rational design of bisubstrate analogue inhibitors of basophilic protein kinases. J Med Chem52:308-21 (2009) [PubMed] Article