| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Smoothened homolog |
|---|
| Ligand | BDBM50399540 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEBML_1690630 |
|---|
| Ki | 60±n/a nM |
|---|
| Citation |  Morgillo, F; Amendola, G; Della Corte, CM; Giacomelli, C; Botta, L; Di Maro, S; Messere, A; Ciaramella, V; Taliani, S; Marinelli, L; Trincavelli, ML; Martini, C; Novellino, E; Ciardiello, F; Cosconati, S Dual MET and SMO Negative Modulators Overcome Resistance to EGFR Inhibitors in Human Nonsmall Cell Lung Cancer. J Med Chem60:7447-7458 (2017) [PubMed] Article Morgillo, F; Amendola, G; Della Corte, CM; Giacomelli, C; Botta, L; Di Maro, S; Messere, A; Ciaramella, V; Taliani, S; Marinelli, L; Trincavelli, ML; Martini, C; Novellino, E; Ciardiello, F; Cosconati, S Dual MET and SMO Negative Modulators Overcome Resistance to EGFR Inhibitors in Human Nonsmall Cell Lung Cancer. J Med Chem60:7447-7458 (2017) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Smoothened homolog |
|---|
| Name: | Smoothened homolog |
|---|
| Synonyms: | G-protein- coupled-like receptor Smoothened (Smo) | SMO | SMOH | SMO_HUMAN | Smoothened homolog |
|---|
| Type: | Protein |
|---|
| Mol. Mass.: | 86415.51 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | Q99835 |
|---|
| Residue: | 787 |
|---|
| Sequence: | MAAARPARGPELPLLGLLLLLLLGDPGRGAASSGNATGPGPRSAGGSARRSAAVTGPPPP
LSHCGRAAPCEPLRYNVCLGSVLPYGATSTLLAGDSDSQEEAHGKLVLWSGLRNAPRCWA
VIQPLLCAVYMPKCENDRVELPSRTLCQATRGPCAIVERERGWPDFLRCTPDRFPEGCTN
EVQNIKFNSSGQCEVPLVRTDNPKSWYEDVEGCGIQCQNPLFTEAEHQDMHSYIAAFGAV
TGLCTLFTLATFVADWRNSNRYPAVILFYVNACFFVGSIGWLAQFMDGARREIVCRADGT
MRLGEPTSNETLSCVIIFVIVYYALMAGVVWFVVLTYAWHTSFKALGTTYQPLSGKTSYF
HLLTWSLPFVLTVAILAVAQVDGDSVSGICFVGYKNYRYRAGFVLAPIGLVLIVGGYFLI
RGVMTLFSIKSNHPGLLSEKAASKINETMLRLGIFGFLAFGFVLITFSCHFYDFFNQAEW
ERSFRDYVLCQANVTIGLPTKQPIPDCEIKNRPSLLVEKINLFAMFGTGIAMSTWVWTKA
TLLIWRRTWCRLTGQSDDEPKRIKKSKMIAKAFSKRHELLQNPGQELSFSMHTVSHDGPV
AGLAFDLNEPSADVSSAWAQHVTKMVARRGAILPQDISVTPVATPVPPEEQANLWLVEAE
ISPELQKRLGRKKKRRKRKKEVCPLAPPPELHPPAPAPSTIPRLPQLPRQKCLVAAGAWG
AGDSCRQGAWTLVSNPFCPEPSPPQDPFLPSAPAPVAWAHGRRQGLGPIHSRTNLMDTEL
MDADSDF
|
|
|
|---|
| BDBM50399540 |
|---|
| n/a |
|---|
| Name | BDBM50399540 |
|---|
| Synonyms: | FORETINIB | US10464902, Foretinib | US10882853, Compound For-Oxide |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C34H34F2N4O6 |
|---|
| Mol. Mass. | 632.6538 |
|---|
| SMILES | COc1cc2c(Oc3ccc(NC(=O)C4(CC4)C(=O)Nc4ccc(F)cc4)cc3F)ccnc2cc1OCCCN1CCOCC1 |
|---|
| Structure | 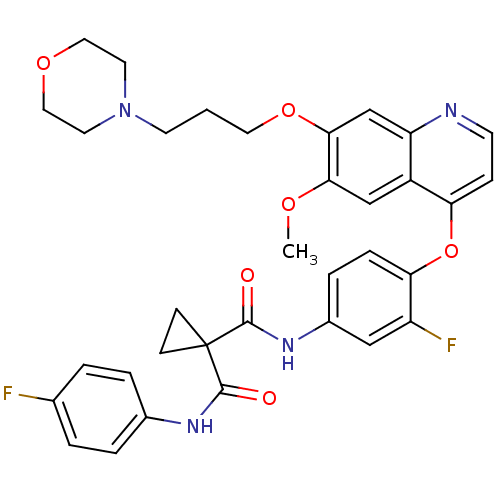
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Morgillo, F; Amendola, G; Della Corte, CM; Giacomelli, C; Botta, L; Di Maro, S; Messere, A; Ciaramella, V; Taliani, S; Marinelli, L; Trincavelli, ML; Martini, C; Novellino, E; Ciardiello, F; Cosconati, S Dual MET and SMO Negative Modulators Overcome Resistance to EGFR Inhibitors in Human Nonsmall Cell Lung Cancer. J Med Chem60:7447-7458 (2017) [PubMed] Article
Morgillo, F; Amendola, G; Della Corte, CM; Giacomelli, C; Botta, L; Di Maro, S; Messere, A; Ciaramella, V; Taliani, S; Marinelli, L; Trincavelli, ML; Martini, C; Novellino, E; Ciardiello, F; Cosconati, S Dual MET and SMO Negative Modulators Overcome Resistance to EGFR Inhibitors in Human Nonsmall Cell Lung Cancer. J Med Chem60:7447-7458 (2017) [PubMed] Article