| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Indoleamine 2,3-dioxygenase 1 |
|---|
| Ligand | BDBM50520449 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_1880241 (CHEMBL4381635) |
|---|
| IC50 | 42±n/a nM |
|---|
| Citation |  Peng, YH; Liao, FY; Tseng, CT; Kuppusamy, R; Li, AS; Chen, CH; Fan, YS; Wang, SY; Wu, MH; Hsueh, CC; Chang, JY; Lee, LC; Shih, C; Shia, KS; Yeh, TK; Hung, MS; Kuo, CC; Song, JS; Wu, SY; Ueng, SH Unique Sulfur-Aromatic Interactions Contribute to the Binding of Potent Imidazothiazole Indoleamine 2,3-Dioxygenase Inhibitors. J Med Chem63:1642-1659 (2020) [PubMed] Article Peng, YH; Liao, FY; Tseng, CT; Kuppusamy, R; Li, AS; Chen, CH; Fan, YS; Wang, SY; Wu, MH; Hsueh, CC; Chang, JY; Lee, LC; Shih, C; Shia, KS; Yeh, TK; Hung, MS; Kuo, CC; Song, JS; Wu, SY; Ueng, SH Unique Sulfur-Aromatic Interactions Contribute to the Binding of Potent Imidazothiazole Indoleamine 2,3-Dioxygenase Inhibitors. J Med Chem63:1642-1659 (2020) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Indoleamine 2,3-dioxygenase 1 |
|---|
| Name: | Indoleamine 2,3-dioxygenase 1 |
|---|
| Synonyms: | I23O1_HUMAN | IDO | IDO-1 | IDO1 | INDO | Indoleamine 2,3-Dioxygenasae (IDO) | Indoleamine 2,3-dioxygenase | Indoleamine-pyrrole 2,3-dioxygenase |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 45330.80 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | P14902 |
|---|
| Residue: | 403 |
|---|
| Sequence: | MAHAMENSWTISKEYHIDEEVGFALPNPQENLPDFYNDWMFIAKHLPDLIESGQLRERVE
KLNMLSIDHLTDHKSQRLARLVLGCITMAYVWGKGHGDVRKVLPRNIAVPYCQLSKKLEL
PPILVYADCVLANWKKKDPNKPLTYENMDVLFSFRDGDCSKGFFLVSLLVEIAAASAIKV
IPTVFKAMQMQERDTLLKALLEIASCLEKALQVFHQIHDHVNPKAFFSVLRIYLSGWKGN
PQLSDGLVYEGFWEDPKEFAGGSAGQSSVFQCFDVLLGIQQTAGGGHAAQFLQDMRRYMP
PAHRNFLCSLESNPSVREFVLSKGDAGLREAYDACVKALVSLRSYHLQIVTKYILIPASQ
QPKENKTSEDPSKLEAKGTGGTDLMNFLKTVRSTTEKSLLKEG
|
|
|
|---|
| BDBM50520449 |
|---|
| n/a |
|---|
| Name | BDBM50520449 |
|---|
| Synonyms: | CHEMBL4459210 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C18H15ClN4OS |
|---|
| Mol. Mass. | 370.856 |
|---|
| SMILES | Clc1cccc(NC(=O)NCc2cnc3scc(C#CC4CC4)n23)c1 |
|---|
| Structure | 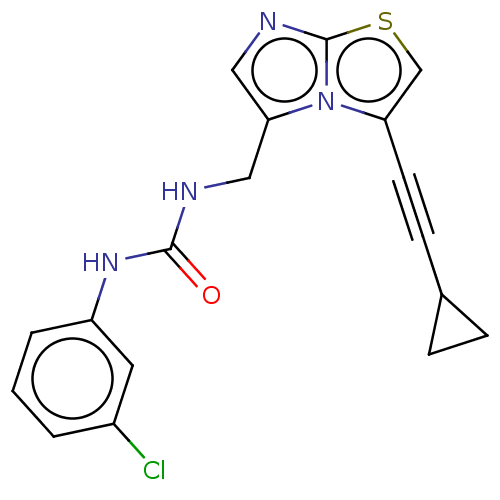
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Peng, YH; Liao, FY; Tseng, CT; Kuppusamy, R; Li, AS; Chen, CH; Fan, YS; Wang, SY; Wu, MH; Hsueh, CC; Chang, JY; Lee, LC; Shih, C; Shia, KS; Yeh, TK; Hung, MS; Kuo, CC; Song, JS; Wu, SY; Ueng, SH Unique Sulfur-Aromatic Interactions Contribute to the Binding of Potent Imidazothiazole Indoleamine 2,3-Dioxygenase Inhibitors. J Med Chem63:1642-1659 (2020) [PubMed] Article
Peng, YH; Liao, FY; Tseng, CT; Kuppusamy, R; Li, AS; Chen, CH; Fan, YS; Wang, SY; Wu, MH; Hsueh, CC; Chang, JY; Lee, LC; Shih, C; Shia, KS; Yeh, TK; Hung, MS; Kuo, CC; Song, JS; Wu, SY; Ueng, SH Unique Sulfur-Aromatic Interactions Contribute to the Binding of Potent Imidazothiazole Indoleamine 2,3-Dioxygenase Inhibitors. J Med Chem63:1642-1659 (2020) [PubMed] Article