| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Prolyl endopeptidase FAP |
|---|
| Ligand | BDBM50171555 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_321439 (CHEMBL880247) |
|---|
| IC50 | 6800±n/a nM |
|---|
| Citation |  Shreder, KR; Wong, MS; Corral, S; Yu, Z; Winn, DT; Wu, M; Hu, Y; Nomanbhoy, T; Alemayehu, S; Fuller, SR; Rosenblum, JS; Kozarich, JW Boro-norleucine as a P1 residue for the design of selective and potent DPP7 inhibitors. Bioorg Med Chem Lett15:4256-60 (2005) [PubMed] Article Shreder, KR; Wong, MS; Corral, S; Yu, Z; Winn, DT; Wu, M; Hu, Y; Nomanbhoy, T; Alemayehu, S; Fuller, SR; Rosenblum, JS; Kozarich, JW Boro-norleucine as a P1 residue for the design of selective and potent DPP7 inhibitors. Bioorg Med Chem Lett15:4256-60 (2005) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Prolyl endopeptidase FAP |
|---|
| Name: | Prolyl endopeptidase FAP |
|---|
| Synonyms: | 170 kDa melanoma membrane-bound gelatinase | FAP | Fibroblast Activation Protein (FAP) | Fibroblast activation protein alpha | Integral membrane serine protease | SEPR_HUMAN | Seprase |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 87712.48 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | Q12884 |
|---|
| Residue: | 760 |
|---|
| Sequence: | MKTWVKIVFGVATSAVLALLVMCIVLRPSRVHNSEENTMRALTLKDILNGTFSYKTFFPN
WISGQEYLHQSADNNIVLYNIETGQSYTILSNRTMKSVNASNYGLSPDRQFVYLESDYSK
LWRYSYTATYYIYDLSNGEFVRGNELPRPIQYLCWSPVGSKLAYVYQNNIYLKQRPGDPP
FQITFNGRENKIFNGIPDWVYEEEMLATKYALWWSPNGKFLAYAEFNDTDIPVIAYSYYG
DEQYPRTINIPYPKAGAKNPVVRIFIIDTTYPAYVGPQEVPVPAMIASSDYYFSWLTWVT
DERVCLQWLKRVQNVSVLSICDFREDWQTWDCPKTQEHIEESRTGWAGGFFVSTPVFSYD
AISYYKIFSDKDGYKHIHYIKDTVENAIQITSGKWEAINIFRVTQDSLFYSSNEFEEYPG
RRNIYRISIGSYPPSKKCVTCHLRKERCQYYTASFSDYAKYYALVCYGPGIPISTLHDGR
TDQEIKILEENKELENALKNIQLPKEEIKKLEVDEITLWYKMILPPQFDRSKKYPLLIQV
YGGPCSQSVRSVFAVNWISYLASKEGMVIALVDGRGTAFQGDKLLYAVYRKLGVYEVEDQ
ITAVRKFIEMGFIDEKRIAIWGWSYGGYVSSLALASGTGLFKCGIAVAPVSSWEYYASVY
TERFMGLPTKDDNLEHYKNSTVMARAEYFRNVDYLLIHGTADDNVHFQNSAQIAKALVNA
QVDFQAMWYSDQNHGLSGLSTNHLYTHMTHFLKQCFSLSD
|
|
|
|---|
| BDBM50171555 |
|---|
| n/a |
|---|
| Name | BDBM50171555 |
|---|
| Synonyms: | (S)-2,4-Diamino-1-(2-boron-dihydroxide-pyrrolidin-1-yl)-butan-1-one | CHEMBL194428 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C8H18BN3O3 |
|---|
| Mol. Mass. | 215.058 |
|---|
| SMILES | NCC[C@H](N)C(=O)N1CCC[C@@H]1B(O)O |
|---|
| Structure | 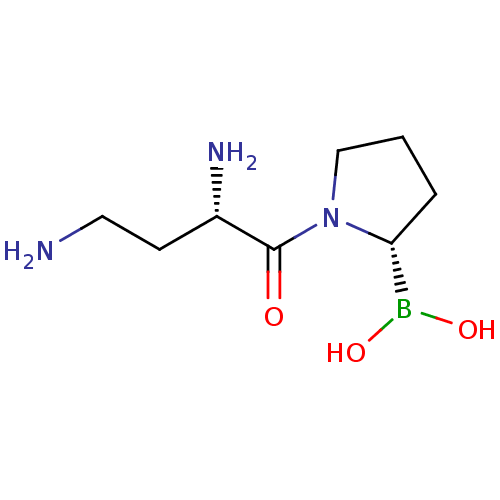
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Shreder, KR; Wong, MS; Corral, S; Yu, Z; Winn, DT; Wu, M; Hu, Y; Nomanbhoy, T; Alemayehu, S; Fuller, SR; Rosenblum, JS; Kozarich, JW Boro-norleucine as a P1 residue for the design of selective and potent DPP7 inhibitors. Bioorg Med Chem Lett15:4256-60 (2005) [PubMed] Article
Shreder, KR; Wong, MS; Corral, S; Yu, Z; Winn, DT; Wu, M; Hu, Y; Nomanbhoy, T; Alemayehu, S; Fuller, SR; Rosenblum, JS; Kozarich, JW Boro-norleucine as a P1 residue for the design of selective and potent DPP7 inhibitors. Bioorg Med Chem Lett15:4256-60 (2005) [PubMed] Article