| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Glutamyl aminopeptidase |
|---|
| Ligand | BDBM50083384 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_32830 (CHEMBL873060) |
|---|
| Ki | 20±n/a nM |
|---|
| Citation |  David, C; Bischoff, L; Meudal, H; Mothé, A; De Mota, N; DaNascimento, S; Llorens-Cortes, C; Fournié-Zaluski, MC; Roques, BP Investigation of subsite preferences in aminopeptidase A (EC 3.4.11.7) led to the design of the first highly potent and selective inhibitors of this enzyme. J Med Chem42:5197-211 (2000) [PubMed] David, C; Bischoff, L; Meudal, H; Mothé, A; De Mota, N; DaNascimento, S; Llorens-Cortes, C; Fournié-Zaluski, MC; Roques, BP Investigation of subsite preferences in aminopeptidase A (EC 3.4.11.7) led to the design of the first highly potent and selective inhibitors of this enzyme. J Med Chem42:5197-211 (2000) [PubMed] |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Glutamyl aminopeptidase |
|---|
| Name: | Glutamyl aminopeptidase |
|---|
| Synonyms: | AMPE_HUMAN | AP-A | Aminopeptidase A | CD_antigen=CD249 | Differentiation antigen gp160 | EAP | ENPEP | Glutamyl aminopeptidase |
|---|
| Type: | PROTEIN |
|---|
| Mol. Mass.: | 109226.47 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | ChEMBL_32829 |
|---|
| Residue: | 957 |
|---|
| Sequence: | MNFAEREGSKRYCIQTKHVAILCAVVVGVGLIVGLAVGLTRSCDSSGDGGPGTAPAPSHL
PSSTASPSGPPAQDQDICPASEDESGQWKNFRLPDFVNPVHYDLHVKPLLEEDTYTGTVS
ISINLSAPTRYLWLHLRETRITRLPELKRPSGDQVQVRRCFEYKKQEYVVVEAEEELTPS
SGDGLYLLTMEFAGWLNGSLVGFYRTTYTENGQVKSIVATDHEPTDARKSFPCFDEPNKK
ATYTISITHPKEYGALSNMPVAKEESVDDKWTRTTFEKSVPMSTYLVCFAVHQFDSVKRI
SNSGKPLTIYVQPEQKHTAEYAANITKSVFDYFEEYFAMNYSLPKLDKIAIPDFGTGAME
NWGLITYRETNLLYDPKESASSNQQRVATVVAHELVHQWFGNIVTMDWWEDLWLNEGFAS
FFEFLGVNHAETDWQMRDQMLLEDVLPVQEDDSLMSSHPIIVTVTTPDEITSVFDGISYS
KGSSILRMLEDWIKPENFQKGCQMYLEKYQFKNAKTSDFWAALEEASRLPVKEVMDTWTR
QMGYPVLNVNGVKNITQKRFLLDPRANPSQPPSDLGYTWNIPVKWTEDNITSSVLFNRSE
KEGITLNSSNPSGNAFLKINPDHIGFYRVNYEVATWDSIATALSLNHKTFSSADRASLID
DAFALARAQLLDYKVALNLTKYLKREENFLPWQRVISAVTYIISMFEDDKELYPMIEEYF
QGQVKPIADSLGWNDAGDHVTKLLRSSVLGFACKMGDREALNNASSLFEQWLNGTVSLPV
NLRLLVYRYGMQNSGNEISWNYTLEQYQKTSLAQEKEKLLYGLASVKNVTLLSRYLDLLK
DTNLIKTQDVFTVIRYISYNSYGKNMAWNWIQLNWDYLVNRYTLNNRNLGRIVTIAEPFN
TELQLWQMESFFAKYPQAGAGEKPREQVLETVKNNIEWLKQHRNTIREWFFNLLESG
|
|
|
|---|
| BDBM50083384 |
|---|
| n/a |
|---|
| Name | BDBM50083384 |
|---|
| Synonyms: | 1-{[1-(1,2-Dicarboxy-ethylcarbamoyl)-2-methyl-butylcarbamoyl]-mercapto-methyl}-3-sulfo-propyl-ammonium | CHEMBL345941 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C15H27N3O9S2 |
|---|
| Mol. Mass. | 457.52 |
|---|
| SMILES | CC[C@@H](C)[C@H](NC(=O)[C@H](S)[C@@H]([NH3+])CCS([O-])(=O)=O)C(=O)N[C@@H](CC(O)=O)C(O)=O |
|---|
| Structure | 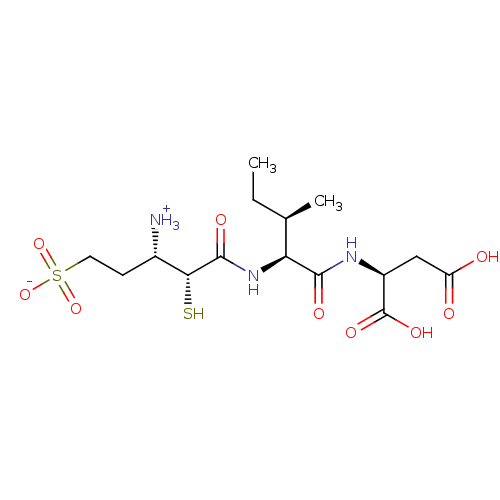
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 David, C; Bischoff, L; Meudal, H; Mothé, A; De Mota, N; DaNascimento, S; Llorens-Cortes, C; Fournié-Zaluski, MC; Roques, BP Investigation of subsite preferences in aminopeptidase A (EC 3.4.11.7) led to the design of the first highly potent and selective inhibitors of this enzyme. J Med Chem42:5197-211 (2000) [PubMed]
David, C; Bischoff, L; Meudal, H; Mothé, A; De Mota, N; DaNascimento, S; Llorens-Cortes, C; Fournié-Zaluski, MC; Roques, BP Investigation of subsite preferences in aminopeptidase A (EC 3.4.11.7) led to the design of the first highly potent and selective inhibitors of this enzyme. J Med Chem42:5197-211 (2000) [PubMed]