| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Methionine aminopeptidase |
|---|
| Ligand | BDBM50129640 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_105132 (CHEMBL872687) |
|---|
| IC50 | 220±n/a nM |
|---|
| Citation |  Luo, QL; Li, JY; Liu, ZY; Chen, LL; Li, J; Qian, Z; Shen, Q; Li, Y; Lushington, GH; Ye, QZ; Nan, FJ Discovery and structural modification of inhibitors of methionine aminopeptidases from Escherichia coli and Saccharomyces cerevisiae. J Med Chem46:2631-40 (2003) [PubMed] Article Luo, QL; Li, JY; Liu, ZY; Chen, LL; Li, J; Qian, Z; Shen, Q; Li, Y; Lushington, GH; Ye, QZ; Nan, FJ Discovery and structural modification of inhibitors of methionine aminopeptidases from Escherichia coli and Saccharomyces cerevisiae. J Med Chem46:2631-40 (2003) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Methionine aminopeptidase |
|---|
| Name: | Methionine aminopeptidase |
|---|
| Synonyms: | EcMetAP | MAP1_ECOLI | Methionine Aminopeptidase (MAP) | Methionine aminopeptidase | Peptidase M | map |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 29326.96 |
|---|
| Organism: | Escherichia coli (strain K12) |
|---|
| Description: | Full-length untagged EcMAP was expressed in E. coli. |
|---|
| Residue: | 264 |
|---|
| Sequence: | MAISIKTPEDIEKMRVAGRLAAEVLEMIEPYVKPGVSTGELDRICNDYIVNEQHAVSACL
GYHGYPKSVCISINEVVCHGIPDDAKLLKDGDIVNIDVTVIKDGFHGDTSKMFIVGKPTI
MGERLCRITQESLYLALRMVKPGINLREIGAAIQKFVEAEGFSVVREYCGHGIGRGFHEE
PQVLHYDSRETNVVLKPGMTFTIEPMVNAGKKEIRTMKDGWTVKTKDRSLSAQYEHTIVV
TDNGCEILTLRKDDTIPAIISHDE
|
|
|
|---|
| BDBM50129640 |
|---|
| n/a |
|---|
| Name | BDBM50129640 |
|---|
| Synonyms: | 3-(3-Methyl-but-2-enoylamino)-pyridine-2-carboxylic acid thiazol-2-ylamide | CHEMBL313629 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C14H14N4O2S |
|---|
| Mol. Mass. | 302.352 |
|---|
| SMILES | [#6]\[#6](-[#6])=[#6]\[#6](=O)-[#7]-c1cccnc1-[#6](=O)-[#7]-c1nccs1 |
|---|
| Structure | 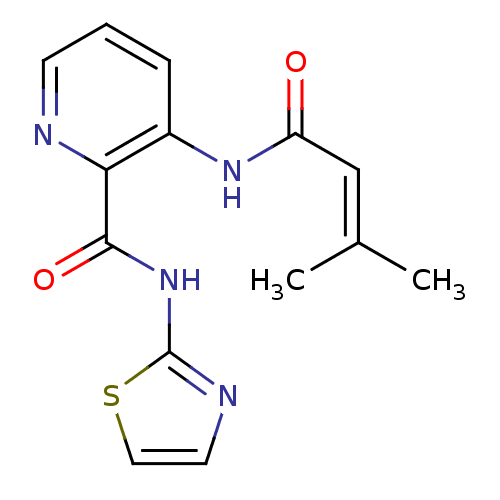
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Luo, QL; Li, JY; Liu, ZY; Chen, LL; Li, J; Qian, Z; Shen, Q; Li, Y; Lushington, GH; Ye, QZ; Nan, FJ Discovery and structural modification of inhibitors of methionine aminopeptidases from Escherichia coli and Saccharomyces cerevisiae. J Med Chem46:2631-40 (2003) [PubMed] Article
Luo, QL; Li, JY; Liu, ZY; Chen, LL; Li, J; Qian, Z; Shen, Q; Li, Y; Lushington, GH; Ye, QZ; Nan, FJ Discovery and structural modification of inhibitors of methionine aminopeptidases from Escherichia coli and Saccharomyces cerevisiae. J Med Chem46:2631-40 (2003) [PubMed] Article