| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Peroxisome proliferator-activated receptor gamma |
|---|
| Ligand | BDBM50372111 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_464735 (CHEMBL933081) |
|---|
| EC50 | 60±n/a nM |
|---|
| Citation |  Wang, W; Devasthale, P; Farrelly, D; Gu, L; Harrity, T; Cap, M; Chu, C; Kunselman, L; Morgan, N; Ponticiello, R; Zebo, R; Zhang, L; Locke, K; Lippy, J; O'Malley, K; Hosagrahara, V; Zhang, L; Kadiyala, P; Chang, C; Muckelbauer, J; Doweyko, AM; Zahler, R; Ryono, D; Hariharan, N; Cheng, PT Discovery of azetidinone acids as conformationally-constrained dual PPARalpha/gamma agonists. Bioorg Med Chem Lett18:1939-44 (2008) [PubMed] Article Wang, W; Devasthale, P; Farrelly, D; Gu, L; Harrity, T; Cap, M; Chu, C; Kunselman, L; Morgan, N; Ponticiello, R; Zebo, R; Zhang, L; Locke, K; Lippy, J; O'Malley, K; Hosagrahara, V; Zhang, L; Kadiyala, P; Chang, C; Muckelbauer, J; Doweyko, AM; Zahler, R; Ryono, D; Hariharan, N; Cheng, PT Discovery of azetidinone acids as conformationally-constrained dual PPARalpha/gamma agonists. Bioorg Med Chem Lett18:1939-44 (2008) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Peroxisome proliferator-activated receptor gamma |
|---|
| Name: | Peroxisome proliferator-activated receptor gamma |
|---|
| Synonyms: | NR1C3 | Nuclear receptor subfamily 1 group C member 3 | PPAR-gamma | PPARG | PPARG_HUMAN | Peroxisome proliferator-activated receptor | Peroxisome proliferator-activated receptor gamma (PPAR gamma) | Peroxisome proliferator-activated receptor gamma (PPARG) | Peroxisome proliferator-activated receptor gamma (PPARγ) | Peroxisome proliferator-activated receptor gamma/Nuclear receptor corepressor 2 | peroxisome proliferator-activated receptor gamma isoform 2 |
|---|
| Type: | Nuclear Receptor |
|---|
| Mol. Mass.: | 57613.46 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | P37231 |
|---|
| Residue: | 505 |
|---|
| Sequence: | MGETLGDSPIDPESDSFTDTLSANISQEMTMVDTEMPFWPTNFGISSVDLSVMEDHSHSF
DIKPFTTVDFSSISTPHYEDIPFTRTDPVVADYKYDLKLQEYQSAIKVEPASPPYYSEKT
QLYNKPHEEPSNSLMAIECRVCGDKASGFHYGVHACEGCKGFFRRTIRLKLIYDRCDLNC
RIHKKSRNKCQYCRFQKCLAVGMSHNAIRFGRMPQAEKEKLLAEISSDIDQLNPESADLR
ALAKHLYDSYIKSFPLTKAKARAILTGKTTDKSPFVIYDMNSLMMGEDKIKFKHITPLQE
QSKEVAIRIFQGCQFRSVEAVQEITEYAKSIPGFVNLDLNDQVTLLKYGVHEIIYTMLAS
LMNKDGVLISEGQGFMTREFLKSLRKPFGDFMEPKFEFAVKFNALELDDSDLAIFIAVII
LSGDRPGLLNVKPIEDIQDNLLQALELQLKLNHPESSQLFAKLLQKMTDLRQIVTEHVQL
LQVIKKTETDMSLHPLLQEIYKDLY
|
|
|
|---|
| BDBM50372111 |
|---|
| n/a |
|---|
| Name | BDBM50372111 |
|---|
| Synonyms: | CHEMBL272336 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C33H33ClN2O5 |
|---|
| Mol. Mass. | 573.079 |
|---|
| SMILES | Cc1oc(nc1CCOc1cccc(C[C@H]2[C@H](N(C2=O)c2ccc(cc2)C(C)(C)C)C(O)=O)c1)-c1ccc(Cl)cc1 |
|---|
| Structure | 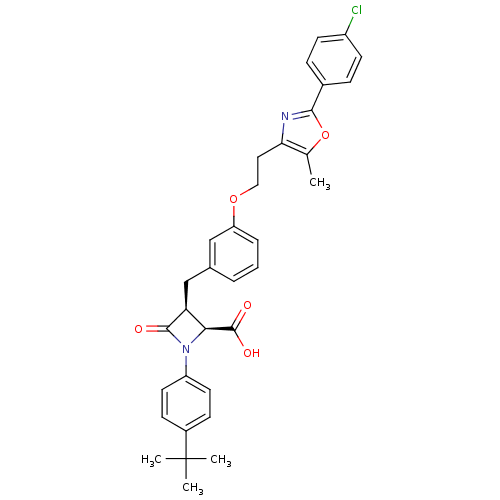
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Wang, W; Devasthale, P; Farrelly, D; Gu, L; Harrity, T; Cap, M; Chu, C; Kunselman, L; Morgan, N; Ponticiello, R; Zebo, R; Zhang, L; Locke, K; Lippy, J; O'Malley, K; Hosagrahara, V; Zhang, L; Kadiyala, P; Chang, C; Muckelbauer, J; Doweyko, AM; Zahler, R; Ryono, D; Hariharan, N; Cheng, PT Discovery of azetidinone acids as conformationally-constrained dual PPARalpha/gamma agonists. Bioorg Med Chem Lett18:1939-44 (2008) [PubMed] Article
Wang, W; Devasthale, P; Farrelly, D; Gu, L; Harrity, T; Cap, M; Chu, C; Kunselman, L; Morgan, N; Ponticiello, R; Zebo, R; Zhang, L; Locke, K; Lippy, J; O'Malley, K; Hosagrahara, V; Zhang, L; Kadiyala, P; Chang, C; Muckelbauer, J; Doweyko, AM; Zahler, R; Ryono, D; Hariharan, N; Cheng, PT Discovery of azetidinone acids as conformationally-constrained dual PPARalpha/gamma agonists. Bioorg Med Chem Lett18:1939-44 (2008) [PubMed] Article