| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Peroxisome proliferator-activated receptor delta |
|---|
| Ligand | BDBM50127222 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_718368 (CHEMBL1679900) |
|---|
| EC50 | 300±n/a nM |
|---|
| Citation |  Patch, RJ; Searle, LL; Kim, AJ; De, D; Zhu, X; Askari, HB; O'Neill, JC; Abad, MC; Rentzeperis, D; Liu, J; Kemmerer, M; Lin, L; Kasturi, J; Geisler, JG; Lenhard, JM; Player, MR; Gaul, MD Identification of diaryl ether-based ligands for estrogen-related receptora as potential antidiabetic agents. J Med Chem54:788-808 (2012) [PubMed] Article Patch, RJ; Searle, LL; Kim, AJ; De, D; Zhu, X; Askari, HB; O'Neill, JC; Abad, MC; Rentzeperis, D; Liu, J; Kemmerer, M; Lin, L; Kasturi, J; Geisler, JG; Lenhard, JM; Player, MR; Gaul, MD Identification of diaryl ether-based ligands for estrogen-related receptora as potential antidiabetic agents. J Med Chem54:788-808 (2012) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Peroxisome proliferator-activated receptor delta |
|---|
| Name: | Peroxisome proliferator-activated receptor delta |
|---|
| Synonyms: | NR1C2 | NUC1 | NUCI | Nuclear hormone receptor 1 | Nuclear receptor subfamily 1 group C member 2 | PPAR delta | PPAR-beta | PPARB | PPARD | PPARD_HUMAN | Peroxisome proliferator-activated receptor | Peroxisome proliferator-activated receptor beta | Peroxisome proliferator-activated receptor delta |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 49910.45 |
|---|
| Organism: | Homo sapiens (Human) |
|---|
| Description: | Q03181 |
|---|
| Residue: | 441 |
|---|
| Sequence: | MEQPQEEAPEVREEEEKEEVAEAEGAPELNGGPQHALPSSSYTDLSRSSSPPSLLDQLQM
GCDGASCGSLNMECRVCGDKASGFHYGVHACEGCKGFFRRTIRMKLEYEKCERSCKIQKK
NRNKCQYCRFQKCLALGMSHNAIRFGRMPEAEKRKLVAGLTANEGSQYNPQVADLKAFSK
HIYNAYLKNFNMTKKKARSILTGKASHTAPFVIHDIETLWQAEKGLVWKQLVNGLPPYKE
ISVHVFYRCQCTTVETVRELTEFAKSIPSFSSLFLNDQVTLLKYGVHEAIFAMLASIVNK
DGLLVANGSGFVTREFLRSLRKPFSDIIEPKFEFAVKFNALELDDSDLALFIAAIILCGD
RPGLMNVPRVEAIQDTILRALEFHLQANHPDAQYLFPKLLQKMADLRQLVTEHAQMMQRI
KKTETETSLHPLLQEIYKDMY
|
|
|
|---|
| BDBM50127222 |
|---|
| n/a |
|---|
| Name | BDBM50127222 |
|---|
| Synonyms: | CHEMBL38508 | GW-0742 | {4-[2-(3-Fluoro-4-trifluoromethyl-phenyl)-4-methyl-thiazol-5-ylmethylsulfanyl]-2-methyl-phenoxy}-acetic acid |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C21H17F4NO3S2 |
|---|
| Mol. Mass. | 471.488 |
|---|
| SMILES | Cc1nc(sc1CSc1ccc(OCC(O)=O)c(C)c1)-c1ccc(c(F)c1)C(F)(F)F |
|---|
| Structure | 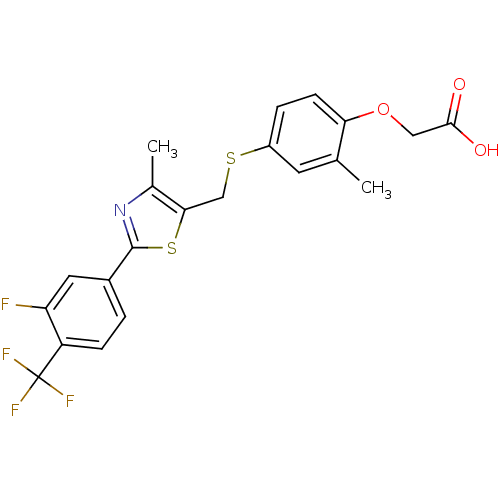
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Patch, RJ; Searle, LL; Kim, AJ; De, D; Zhu, X; Askari, HB; O'Neill, JC; Abad, MC; Rentzeperis, D; Liu, J; Kemmerer, M; Lin, L; Kasturi, J; Geisler, JG; Lenhard, JM; Player, MR; Gaul, MD Identification of diaryl ether-based ligands for estrogen-related receptora as potential antidiabetic agents. J Med Chem54:788-808 (2012) [PubMed] Article
Patch, RJ; Searle, LL; Kim, AJ; De, D; Zhu, X; Askari, HB; O'Neill, JC; Abad, MC; Rentzeperis, D; Liu, J; Kemmerer, M; Lin, L; Kasturi, J; Geisler, JG; Lenhard, JM; Player, MR; Gaul, MD Identification of diaryl ether-based ligands for estrogen-related receptora as potential antidiabetic agents. J Med Chem54:788-808 (2012) [PubMed] Article