

 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data
| Reaction Details | |||
|---|---|---|---|
 | Report a problem with these data | ||
| Target | E3 ubiquitin-protein ligase Mdm2 | ||
| Ligand | BDBM162023 | ||
| Substrate/Competitor | n/a | ||
| Meas. Tech. | Time Resolved Fluorescence Energy Transfer (TR-FRET) Assay | ||
| IC50 | 1.9±n/a nM | ||
| Citation |  Berghausen, J; Buschmann, N; Furet, P; Gessier, F; Hergovich Lisztwan, J; Holzer, P; Jacoby, E; Kallen, J; Masuya, K; Pissot Soldermann, C; Ren, H; Stutz, S Substituted isoquinolinones and quinazolinones US Patent US9051279 Publication Date 6/9/2015 Berghausen, J; Buschmann, N; Furet, P; Gessier, F; Hergovich Lisztwan, J; Holzer, P; Jacoby, E; Kallen, J; Masuya, K; Pissot Soldermann, C; Ren, H; Stutz, S Substituted isoquinolinones and quinazolinones US Patent US9051279 Publication Date 6/9/2015 | ||
| More Info.: | Get all data from this article, Assay Method | ||
| E3 ubiquitin-protein ligase Mdm2 | |||
| Name: | E3 ubiquitin-protein ligase Mdm2 | ||
| Synonyms: | Double minute 2 protein | Double minute 2 protein (HDM2) | E3 ubiquitin-protein ligase Mdm2 (p53-binding protein Mdm2) | Hdm2 | Human Double Minute 2 (HDM2) | MDM2 | MDM2-MDMX | MDM2_HUMAN | p53-Binding Protein MDM2 | p53-binding protein | ||
| Type: | Oncoprotein | ||
| Mol. Mass.: | 55196.54 | ||
| Organism: | Homo sapiens (Human) | ||
| Description: | Q00987 | ||
| Residue: | 491 | ||
| Sequence: |
| ||
| BDBM162023 | |||
| n/a | |||
| Name | BDBM162023 | ||
| Synonyms: | US9051279, 73a | US9051279, 73b | ||
| Type | Small organic molecule | ||
| Emp. Form. | C36H46ClN3O3 | ||
| Mol. Mass. | 604.222 | ||
| SMILES | CC[C@@H](C)Oc1cc2[C@@H](N(C(=O)Cc2cc1OC)c1ccc(cc1)N(C)C[C@H]1CC[C@@H](CC1)N(C)C)c1ccc(Cl)cc1 |r,wU:30.36,wD:8.39,2.2,27.29,(9.34,7.63,;8,6.86,;6.67,7.63,;6.67,9.17,;5.33,6.86,;4,7.63,;2.67,6.86,;1.33,7.63,;,6.86,;-1.33,7.63,;-1.33,9.17,;-2.67,9.94,;,9.94,;1.33,9.17,;2.67,9.94,;4,9.17,;5.33,9.94,;5.33,11.48,;-2.67,6.86,;-2.67,5.32,;-4,4.55,;-5.33,5.32,;-5.33,6.86,;-4,7.63,;-6.67,4.55,;-6.67,3.01,;-8,5.32,;-8,6.86,;-6.67,7.63,;-6.67,9.17,;-8,9.94,;-9.34,9.17,;-9.34,7.63,;-8,11.48,;-9.34,12.25,;-6.67,12.25,;,5.32,;-1.33,4.55,;-1.33,3.01,;,2.24,;,.7,;1.33,3.01,;1.33,4.55,)| | ||
| Structure | 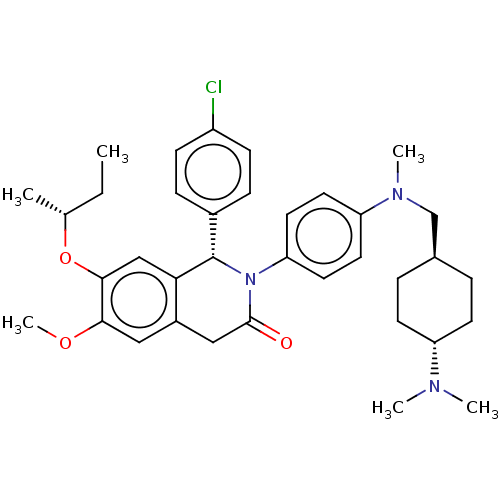
| ||