

 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data
| Reaction Details | |||
|---|---|---|---|
 | Report a problem with these data | ||
| Target | Induced myeloid leukemia cell differentiation protein Mcl-1 [171-327] | ||
| Ligand | BDBM203876 | ||
| Substrate/Competitor | n/a | ||
| Meas. Tech. | TR-FRET Binding A | ||
| pH | 7.4±n/a | ||
| Temperature | 297.15±n/a K | ||
| Ki | 1.6±0 nM | ||
| IC50 | 5.96±0.35 nM | ||
| Comments | extracted | ||
| Citation |  Ak�ay, G; Belmonte, MA; Aquila, B; Chuaqui, C; Hird, AW; Lamb, ML; Rawlins, PB; Su, N; Tentarelli, S; Grimster, NP; Su, Q Inhibition of Mcl-1 through covalent modification of a noncatalytic lysine side chain. Nat Chem Biol12:931-936 (2016) [PubMed] Article Ak�ay, G; Belmonte, MA; Aquila, B; Chuaqui, C; Hird, AW; Lamb, ML; Rawlins, PB; Su, N; Tentarelli, S; Grimster, NP; Su, Q Inhibition of Mcl-1 through covalent modification of a noncatalytic lysine side chain. Nat Chem Biol12:931-936 (2016) [PubMed] Article | ||
| More Info.: | Get all data from this article, Assay Method | ||
| Induced myeloid leukemia cell differentiation protein Mcl-1 [171-327] | |||
| Name: | Induced myeloid leukemia cell differentiation protein Mcl-1 [171-327] | ||
| Synonyms: | BCL2L3 | Induced myeloid leukemia cell differentiation protein Mcl-1(171-327) | MCL1 | MCL1_HUMAN | Myeloid cell leukemia 1 (Mcl-1) | ||
| Type: | Enzyme Catalytic Domain | ||
| Mol. Mass.: | 17896.13 | ||
| Organism: | Homo sapiens (Human) | ||
| Description: | Q07820[171-327] | ||
| Residue: | 157 | ||
| Sequence: |
| ||
| BDBM203876 | |||
| n/a | |||
| Name | BDBM203876 | ||
| Synonyms: | Mcl-1 inhibitor 12 | ||
| Type | Small organic molecule | ||
| Emp. Form. | C36H42N4O4 | ||
| Mol. Mass. | 594.7431 | ||
| SMILES | CCc1nn(C)c(CC)c1-c1cccc2c(CCCOc3cccc4ccccc34)c(C(O)=O)n(CCN3CCOCC3)c12 |(-6.2,-2.88,;-5.11,-1.8,;-3.78,-2.57,;-3.3,-4.03,;-1.76,-4.03,;-.67,-5.12,;-1.28,-2.57,;-.02,-2.6,;.3,-4.24,;-2.53,-1.66,;-2.48,-.6,;-3.81,.17,;-3.81,1.71,;-2.48,2.48,;-1.14,1.71,;.32,2.18,;-.08,3.67,;1.01,4.76,;.62,6.24,;1.7,7.33,;1.31,8.82,;-.03,9.59,;-.03,11.13,;1.31,11.9,;2.64,11.13,;3.97,11.9,;5.31,11.13,;5.31,9.59,;3.97,8.82,;2.64,9.59,;1.23,.94,;2.77,.94,;3.86,-.15,;3.86,2.02,;.32,-.31,;1.22,-1.52,;2.76,-1.52,;3.16,-3.01,;1.82,-3.78,;1.82,-5.32,;3.16,-6.09,;4.49,-5.32,;4.49,-3.78,;-1.14,.17,)| | ||
| Structure | 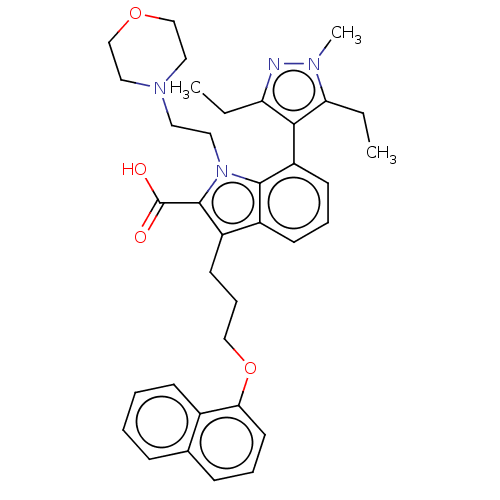
| ||