| Reaction Details |
|---|
 | Report a problem with these data |
| Target | UDP-3-O-acyl-N-acetylglucosamine deacetylase |
|---|
| Ligand | BDBM264723 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_1881663 (CHEMBL4383162) |
|---|
| Kd | 0.440000±n/a nM |
|---|
| Citation |  Lee, PS; Lapointe, G; Madera, AM; Simmons, RL; Xu, W; Yifru, A; Tjandra, M; Karur, S; Rico, A; Thompson, K; Bojkovic, J; Xie, L; Uehara, K; Liu, A; Shu, W; Bellamacina, C; McKenney, D; Morris, L; Tonn, GR; Osborne, C; Benton, BM; McDowell, L; Fu, J; Sweeney, ZK Application of Virtual Screening to the Identification of New LpxC Inhibitor Chemotypes, Oxazolidinone and Isoxazoline. J Med Chem61:9360-9370 (2018) [PubMed] Article Lee, PS; Lapointe, G; Madera, AM; Simmons, RL; Xu, W; Yifru, A; Tjandra, M; Karur, S; Rico, A; Thompson, K; Bojkovic, J; Xie, L; Uehara, K; Liu, A; Shu, W; Bellamacina, C; McKenney, D; Morris, L; Tonn, GR; Osborne, C; Benton, BM; McDowell, L; Fu, J; Sweeney, ZK Application of Virtual Screening to the Identification of New LpxC Inhibitor Chemotypes, Oxazolidinone and Isoxazoline. J Med Chem61:9360-9370 (2018) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| UDP-3-O-acyl-N-acetylglucosamine deacetylase |
|---|
| Name: | UDP-3-O-acyl-N-acetylglucosamine deacetylase |
|---|
| Synonyms: | LPXC_PSEAE | Protein envA | UDP-3-O-[3-hydroxymyristoyl] N-acetylglucosamine deacetylase | UDP-3-O-[3-hydroxymyristoyl] N-acetylglucosamine deacetylase (LpxC | UDP-3-O-[3-hydroxymyristoyl] N-acetylglucosamine deacetylase (LpxC) | UDP-3-O-acyl-GlcNAc deacetylase | UDP-3-O-acyl-GlcNAc deacetylase (LpxC) | envA | lpxC |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 33428.15 |
|---|
| Organism: | Pseudomonas aeruginosa |
|---|
| Description: | P47205 |
|---|
| Residue: | 303 |
|---|
| Sequence: | MIKQRTLKNIIRATGVGLHSGEKVYLTLKPAPVDTGIVFCRTDLDPVVEIPARAENVGET
TMSTTLVKGDVKVDTVEHLLSAMAGLGIDNAYVELSASEVPIMDGSAGPFVFLIQSAGLQ
EQEAAKKFIRIKREVSVEEGDKRAVFVPFDGFKVSFEIDFDHPVFRGRTQQASVDFSSTS
FVKEVSRARTFGFMRDIEYLRSQNLALGGSVENAIVVDENRVLNEDGLRYEDEFVKHKIL
DAIGDLYLLGNSLIGEFRGFKSGHALNNQLLRTLIADKDAWEVVTFEDARTAPISYMRPA
AAV
|
|
|
|---|
| BDBM264723 |
|---|
| n/a |
|---|
| Name | BDBM264723 |
|---|
| Synonyms: | US9718792, 2-A |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C17H20N2O5S |
|---|
| Mol. Mass. | 364.416 |
|---|
| SMILES | CC#Cc1ccc(cc1)C1=NO[C@@H](C[C@](C)(C(=O)NO)S(C)(=O)=O)C1 |r,t:10| |
|---|
| Structure | 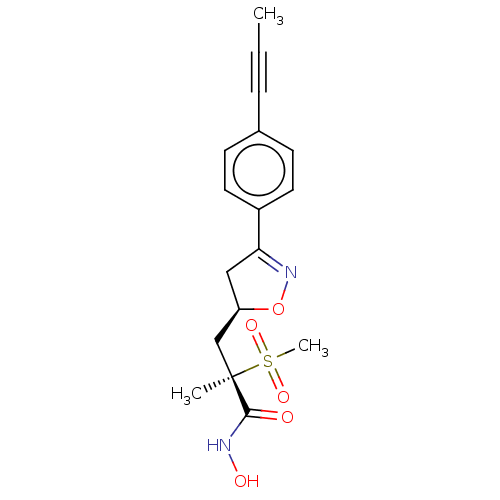
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Lee, PS; Lapointe, G; Madera, AM; Simmons, RL; Xu, W; Yifru, A; Tjandra, M; Karur, S; Rico, A; Thompson, K; Bojkovic, J; Xie, L; Uehara, K; Liu, A; Shu, W; Bellamacina, C; McKenney, D; Morris, L; Tonn, GR; Osborne, C; Benton, BM; McDowell, L; Fu, J; Sweeney, ZK Application of Virtual Screening to the Identification of New LpxC Inhibitor Chemotypes, Oxazolidinone and Isoxazoline. J Med Chem61:9360-9370 (2018) [PubMed] Article
Lee, PS; Lapointe, G; Madera, AM; Simmons, RL; Xu, W; Yifru, A; Tjandra, M; Karur, S; Rico, A; Thompson, K; Bojkovic, J; Xie, L; Uehara, K; Liu, A; Shu, W; Bellamacina, C; McKenney, D; Morris, L; Tonn, GR; Osborne, C; Benton, BM; McDowell, L; Fu, J; Sweeney, ZK Application of Virtual Screening to the Identification of New LpxC Inhibitor Chemotypes, Oxazolidinone and Isoxazoline. J Med Chem61:9360-9370 (2018) [PubMed] Article