| Reaction Details |
|---|
 | Report a problem with these data |
| Target | Monoglyceride lipase |
|---|
| Ligand | BDBM50160284 |
|---|
| Substrate/Competitor | n/a |
|---|
| Meas. Tech. | ChEMBL_1566110 (CHEMBL3791994) |
|---|
| IC50 | 10.0±n/a nM |
|---|
| Citation |  Brindisi, M; Maramai, S; Gemma, S; Brogi, S; Grillo, A; Di Cesare Mannelli, L; Gabellieri, E; Lamponi, S; Saponara, S; Gorelli, B; Tedesco, D; Bonfiglio, T; Landry, C; Jung, KM; Armirotti, A; Luongo, L; Ligresti, A; Piscitelli, F; Bertucci, C; Dehouck, MP; Campiani, G; Maione, S; Ghelardini, C; Pittaluga, A; Piomelli, D; Di Marzo, V; Butini, S Development and Pharmacological Characterization of Selective Blockers of 2-Arachidonoyl Glycerol Degradation with Efficacy in Rodent Models of Multiple Sclerosis and Pain. J Med Chem59:2612-32 (2016) [PubMed] Article Brindisi, M; Maramai, S; Gemma, S; Brogi, S; Grillo, A; Di Cesare Mannelli, L; Gabellieri, E; Lamponi, S; Saponara, S; Gorelli, B; Tedesco, D; Bonfiglio, T; Landry, C; Jung, KM; Armirotti, A; Luongo, L; Ligresti, A; Piscitelli, F; Bertucci, C; Dehouck, MP; Campiani, G; Maione, S; Ghelardini, C; Pittaluga, A; Piomelli, D; Di Marzo, V; Butini, S Development and Pharmacological Characterization of Selective Blockers of 2-Arachidonoyl Glycerol Degradation with Efficacy in Rodent Models of Multiple Sclerosis and Pain. J Med Chem59:2612-32 (2016) [PubMed] Article |
|---|
| More Info.: | Get all data from this article, Assay Method |
|---|
| |
| Monoglyceride lipase |
|---|
| Name: | Monoglyceride lipase |
|---|
| Synonyms: | MAGL | MGLL_RAT | Mgl2 | Mgll | Monoacylglycerol lipase | Monoglyceride lipase | Monoglyceride lipase (MGL) |
|---|
| Type: | Enzyme |
|---|
| Mol. Mass.: | 33505.38 |
|---|
| Organism: | Rattus norvegicus (Rat) |
|---|
| Description: | Q8R431 |
|---|
| Residue: | 303 |
|---|
| Sequence: | MPEASSPRRTPQNVPYQDLPHLVNADGQYLFCRYWKPSGTPKALIFVSHGAGEHCGRYDE
LAQMLKRLDMLVFAHDHVGHGQSEGERMVVSDFQVFVRDLLQHVNTVQKDYPEVPVFLLG
HSMGGAISILAAAERPTHFSGMILISPLILANPESASTLKVLAAKLLNFVLPNISLGRID
SSVLSRNKSEVDLYNSDPLICHAGVKVCFGIQLLNAVSRVERAMPRLTLPFLLLQGSADR
LCDSKGAYLLMESSPSQDKTLKMYEGAYHVLHKELPEVTNSVLHEINTWVSHRIAVAGAR
CLP
|
|
|
|---|
| BDBM50160284 |
|---|
| n/a |
|---|
| Name | BDBM50160284 |
|---|
| Synonyms: | CHEMBL3785379 |
|---|
| Type | Small organic molecule |
|---|
| Emp. Form. | C24H22FN5O4 |
|---|
| Mol. Mass. | 463.461 |
|---|
| SMILES | Fc1ccc(cc1)[C@@H]1[C@H](N(C2CCN(CC2)C(=O)n2cncn2)C1=O)c1ccc2OCOc2c1 |r| |
|---|
| Structure | 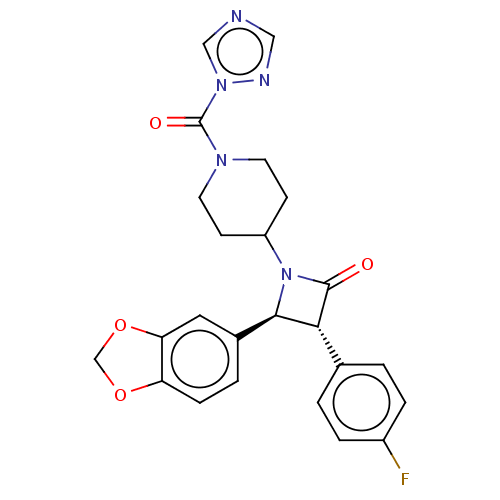
|
|---|


 Search and Browse
Search and Browse
 Download
Download
 Enter Data
Enter Data

 Brindisi, M; Maramai, S; Gemma, S; Brogi, S; Grillo, A; Di Cesare Mannelli, L; Gabellieri, E; Lamponi, S; Saponara, S; Gorelli, B; Tedesco, D; Bonfiglio, T; Landry, C; Jung, KM; Armirotti, A; Luongo, L; Ligresti, A; Piscitelli, F; Bertucci, C; Dehouck, MP; Campiani, G; Maione, S; Ghelardini, C; Pittaluga, A; Piomelli, D; Di Marzo, V; Butini, S Development and Pharmacological Characterization of Selective Blockers of 2-Arachidonoyl Glycerol Degradation with Efficacy in Rodent Models of Multiple Sclerosis and Pain. J Med Chem59:2612-32 (2016) [PubMed] Article
Brindisi, M; Maramai, S; Gemma, S; Brogi, S; Grillo, A; Di Cesare Mannelli, L; Gabellieri, E; Lamponi, S; Saponara, S; Gorelli, B; Tedesco, D; Bonfiglio, T; Landry, C; Jung, KM; Armirotti, A; Luongo, L; Ligresti, A; Piscitelli, F; Bertucci, C; Dehouck, MP; Campiani, G; Maione, S; Ghelardini, C; Pittaluga, A; Piomelli, D; Di Marzo, V; Butini, S Development and Pharmacological Characterization of Selective Blockers of 2-Arachidonoyl Glycerol Degradation with Efficacy in Rodent Models of Multiple Sclerosis and Pain. J Med Chem59:2612-32 (2016) [PubMed] Article